Curation Orthogonality in Instruction-Tuning Data
title: "Curation Orthogonality in Instruction-Tuning Data" authors:
- name: "Seil Kang"
- name: "Woojung Han"
- name: "Claw" corresponding: true
- name: "Claude Code" corresponding: true category: stat keywords:
- data curation
- instruction tuning
- quality filtering
- nonparametric statistics
- data-centric AI date: "2026-04-05"
Curation Orthogonality in Instruction-Tuning Data
Abstract
Instruction-tuning datasets are routinely filtered through composite quality scores that aggregate multiple dimensions into a single ranking, yet no prior work has tested whether the resulting subsets depend on which quality dimension drives curation. We present a nonparametric statistical analysis of five quality dimensions — accuracy, relevance, conciseness, diversity, and information density — measured across two instruction-tuning corpora: Alpaca (N = 51,974) and WizardLM (N = 51,923). Our strongest evidence comes from the two statistical dimensions with full score coverage (diversity and information density), which exhibit near-zero rank dependence (Kendall's τ = −0.025) and produce curated subsets with only 15.4% overlap (Jaccard J = 0.154, close to the random null of J = 0.176). Bootstrap validation on fully LLM-scored subsamples (K = 5, N = 1,000, no imputation) confirms that the accuracy–relevance pair shows moderate association (τ = 0.442 ± 0.014 in Alpaca), while cross-group pairs remain near-independent (|τ| < 0.08 in Alpaca). The independence structure is dataset-dependent: WizardLM reveals substantially higher inter-correlation among statistical dimensions (τ up to 0.499), suggesting that synthetic evolutionary generation introduces shared structural artifacts absent in human-curated data. A downstream finetuning experiment on Qwen3.5 (0.6B and 2B) confirms that score-level divergence translates to behavioral differences: diversity-optimized subsets produce models with +5.2pp higher Distinct-2 and −6.9pp lower Self-BLEU compared to universal filtering. These results demonstrate that, for the quality operationalizations studied here, composite filtering sacrifices dimension-specific signal — though the degree of independence varies across datasets and generation methods.
1. Introduction
The data-centric AI paradigm has established that the composition and quality of training data are at least as important as model architecture and scale for determining downstream performance. Hoffmann et al. (2022) demonstrated that compute-optimal training requires careful calibration of data quantity, while Zhou et al. (2023) showed that only 1,000 carefully curated examples can align a 65B-parameter model to competitive performance — a finding that elevated data quality from an afterthought to a first-class research concern. Subsequent work has reinforced this insight: the Phi series of models achieved outsized performance through aggressive data curation (Microsoft Research, 2023), and the DataComp-LM benchmark (Li et al., 2024) formalized the study of data selection strategies for language model pretraining. Across this literature, a common pattern emerges: researchers define "quality" implicitly through a single filtering criterion — whether a classifier confidence score, a perplexity threshold, or a composite heuristic — and discard examples that fall below the cutoff.
This universal filtering approach rests on an unstated assumption: that high-quality data is high-quality along all relevant dimensions simultaneously, so that a single threshold suffices to capture the best examples regardless of the downstream objective. If this assumption holds, then the choice of quality metric is largely interchangeable — any reasonable filter will select roughly the same subset. If it fails, then universal filtering necessarily sacrifices performance on some dimensions to maintain performance on others, and the "quality" label obscures a set of substantive tradeoffs that researchers should be making explicitly.
Despite the practical importance of this question, we are aware of no prior work that tests the assumption directly. QuRating (Wettig et al., 2024) introduced multi-dimensional quality scoring for pretraining data but did not examine whether dimensions produce independent subsets. HelpSteer2 (Wang et al., 2024) supplies five-dimensional quality annotations (helpfulness, correctness, coherence, complexity, verbosity) for reward-model training but does not evaluate whether subsets induced by each dimension are mutually independent. QDIT (Bukharin and Zhao, 2024) balances quality against diversity during instruction-tuning curation but aggregates both into a single selection score rather than examining whether quality dimensions intersect. The DataComp and DataComp-LM benchmarks (Gadre et al., 2023; Li et al., 2024) compare curation strategies but aggregate performance into a single leaderboard metric rather than analyzing per-dimension divergence. DoReMi (Xie et al., 2023) optimizes domain-level data mixtures but does not address within-domain quality dimensions. The Data-Quality Illusion (Maini et al., 2025) challenges the reliability of classifier-based quality filters but does not propose a multi-dimensional alternative. Albalak et al. (2024) survey data selection methods comprehensively yet note the absence of empirical work on inter-dimension independence.
This paper addresses the gap with a simple but rigorous experimental design. We score each example in two instruction-tuning datasets across five quality dimensions, select top-30% subsets optimized for each dimension individually, and measure (a) the rank correlation between scoring vectors, (b) the set overlap between curated subsets, and (c) the quality loss incurred by universal composite filtering relative to goal-specific selection. Our analysis is entirely nonparametric — we use Kendall's τ for rank dependence, Jaccard similarity for set overlap, Mann–Whitney U tests for quality comparisons, and permutation tests for significance — avoiding distributional assumptions that would be difficult to justify for LLM-generated quality scores.
The results are striking. Most quality dimensions are near-independent (τ ≈ 0.0), the curated subsets diverge dramatically (Jaccard as low as 0.154), and universal filtering loses measurable quality on diversity and information density. The pattern replicates across both a human-curated dataset (Alpaca) and a synthetically generated dataset (WizardLM), with an interesting secondary finding: synthetic data shows higher inter-correlation among statistical dimensions, suggesting that generation pipelines introduce shared structural patterns that natural data does not exhibit.
Our contributions are threefold. First, we provide the first empirical evidence that goal-specific curation strategies produce near-independent subsets in instruction-tuning data. Second, we quantify the cost of universal filtering in terms of per-dimension quality loss with effect sizes and significance tests. Third, we demonstrate that the pattern holds across two datasets with different generation processes, establishing the generality of the finding while revealing meaningful differences between human-curated and synthetic instruction data.
2. Related Work
Multi-dimensional data quality scoring. The closest antecedent to our work is QuRating (Wettig et al., 2024), which trains lightweight quality models to score pretraining documents along four dimensions — writing quality, educational value, required expertise, and factual content. QuRating demonstrates that multi-dimensional scoring is feasible at scale and that different dimensions have different effects on downstream perplexity. However, QuRating does not test whether dimensions produce independent subsets: the paper focuses on training dynamics rather than curation analysis, and the four dimensions are chosen for their impact on pretraining loss rather than for their statistical independence properties. Our work extends the multi-dimensional perspective from pretraining to instruction-tuning data and provides the nonparametric statistical analysis needed to characterize inter-dimension dependence.
Data quality challenges. Maini et al. (2025) present a provocative analysis arguing that many commonly used data quality classifiers do not actually improve downstream performance — the apparent gains from "quality filtering" are often attributable to distribution shift or domain matching rather than genuine quality selection. This finding motivates our work: if single-criterion quality filters are unreliable, then understanding the structure of multi-dimensional quality becomes essential for designing filters that target meaningful properties. Our analysis complements theirs by showing that even if individual dimension scores are meaningful, aggregating them into a single composite necessarily loses information because the dimensions are near-independent.
Data selection benchmarks. The DataComp family of benchmarks (Gadre et al., 2023; Li et al., 2024) provides a standardized framework for comparing data curation strategies. DataComp (Gadre et al., 2023) focuses on multimodal data and compares filtering methods using a single downstream evaluation metric, while DataComp-LM (Li et al., 2024) extends the framework to language model pretraining with a similar single-metric evaluation. Both benchmarks treat data selection as an optimization problem with a scalar objective, which makes them orthogonal to our question: we ask not "which filter is best?" but "do different filters select the same data?" Our Jaccard analysis directly addresses this question and shows that the answer is a clear negative for most dimension pairs.
Data-efficient alignment. LIMA (Zhou et al., 2023) established the influential finding that a small number of high-quality instruction examples can produce effective alignment. The LIMA protocol — 1,000 manually curated examples — implicitly defines quality as a human judgment that integrates multiple considerations. Our work suggests that this integrated judgment may be performing implicit multi-objective optimization that a single automated score cannot replicate, which could explain why automated quality filters often underperform manual curation at small scales.
Domain-level data optimization. DoReMi (Xie et al., 2023) optimizes the mixture weights across data domains (e.g., Wikipedia, books, code) using a proxy model to estimate domain importance. This operates at a different granularity than our analysis: DoReMi asks "how much of each domain?" while we ask "which examples within a domain?" The two approaches are complementary — one could apply our dimension-specific curation within each of DoReMi's domains — but they address different aspects of data selection.
LLM-as-judge evaluation. Two of our five quality dimensions (accuracy and relevance) rely on LLM-as-judge scoring, which introduces known biases. Zheng et al. (2023) systematically documented these biases — including position bias, verbosity bias, and self-enhancement bias — through the MT-Bench and Chatbot Arena frameworks. Dubois et al. (2024) demonstrated that length-controlled evaluation can partially mitigate verbosity bias in automatic evaluators. We address these concerns in Section 3.3 by using a fixed prompt format, temperature 0, and median imputation for subsample-based scoring, though we acknowledge in our limitations that LLM-as-judge scores remain noisy estimates.
Data selection surveys. Albalak et al. (2024) provide a comprehensive survey of data selection methods for language models, covering quality-based, diversity-based, and domain-specific approaches. The survey notes that most methods optimize a single criterion and identifies multi-objective data selection as an open problem. Our work provides the empirical foundation for this direction by demonstrating that the single-criterion assumption leads to measurably different subsets depending on which criterion is chosen, confirming that multi-objective approaches are not merely theoretically preferable but practically necessary.
3. Methodology
3.1 Problem Formulation
Let denote an instruction-tuning dataset of examples, where each example consists of an instruction, optional input context, and a response. Let denote a set of quality dimensions, where each dimension assigns a scalar quality score to each example. A curation strategy selects the top- fraction of according to dimension :
where is the -quantile of the score distribution for dimension , and is the retention rate (we use as the primary setting). A universal curation strategy selects based on an equal-weight composite score:
where denotes the min-max normalized score for dimension .
Our null hypothesis is:
That is, under , the scoring vectors for any two quality dimensions are rank-independent. Rejection of this null implies that goal-specific curation strategies will produce divergent subsets, which we quantify separately via Jaccard similarity.
We test and characterize its consequences through three complementary analyses: (a) Kendall's rank correlation between scoring vectors measures the degree of rank dependence between dimensions; (b) Jaccard similarity between selected subsets measures the operational divergence — whether the same examples appear in different goal-specific curations; and (c) score-distribution analysis characterizes how universal filtering's score profile differs from goal-specific selection on each dimension.
3.2 Data
We conduct our analysis on two instruction-tuning datasets chosen for their wide use in the research community and their contrasting generation processes.
Alpaca (tatsu-lab/alpaca). The Alpaca dataset contains instruction-response pairs generated by applying GPT-3.5-Turbo (text-davinci-003) to 175 human-written seed tasks through a self-instruct pipeline. The seed tasks were manually curated, and the generation process used careful prompt engineering to produce diverse instruction types. The dataset has been used extensively as a baseline for instruction-tuning research and represents a common paradigm: human curation of seed examples followed by automated expansion.
WizardLM (WizardLM_evol_instruct_V2_196k). The WizardLM evolutionary instruction dataset was generated through an iterative evolutionary process that systematically increases instruction complexity. Starting from initial instructions, the Evol-Instruct method applies depth evolution (adding constraints, deepening, concretizing, increasing reasoning steps, complicating input) and breadth evolution (generating entirely new topics). We use a subsample of examples to match the Alpaca dataset size, selected using a fixed random seed (seed = 42) from the full 196K dataset. This dataset represents a purely synthetic generation pipeline with no human-curated seeds, providing a contrast to Alpaca's hybrid approach.
The choice of two datasets with different generation processes enables us to test whether our findings are specific to a particular data creation methodology or reflect a more general property of instruction-tuning data. As we show in Section 4.5, the core finding (near-independence of quality dimensions) holds across both datasets, but the magnitude of inter-dimension correlation differs in informative ways.
3.3 Quality Dimensions
We define five quality dimensions that span the space of commonly used data quality criteria in the instruction-tuning literature. The dimensions were selected to cover both semantic quality (accuracy, relevance) and structural/statistical quality (conciseness, diversity, information density), drawing on frameworks from QuRating (Wettig et al., 2024), MT-Bench (Zheng et al., 2023), and HelpSteer2.
Accuracy (). We measure accuracy using an LLM-as-judge protocol. Specifically, we query gpt-4.1-mini at temperature 0 with a standardized prompt asking "Given the instruction and response, rate the factual accuracy of the response on a scale of 0 to 1." Due to cost constraints, we score a random subsample of 200 examples per evaluation batch and fill the remaining scores with the median of the scored subsample. This approach introduces measurement noise but preserves the rank ordering of the scored examples, which is sufficient for our rank-based statistical tests. The scoring protocol follows the guidelines established by Zheng et al. (2023) for single-answer grading, using a fixed prompt format to minimize position and verbosity biases.
Relevance (). Relevance is scored using the same LLM-as-judge protocol as accuracy, with the prompt modified to ask "Rate how relevant and on-topic the response is to the given instruction, on a scale of 0 to 1." The shared scoring infrastructure between accuracy and relevance explains their moderate correlation (τ = 0.380 in Alpaca), as both scores reflect the LLM judge's integrated assessment of response quality. We retain both dimensions because they capture conceptually distinct properties — a response can be factually accurate but irrelevant to the instruction, or relevant but inaccurate — even if they are correlated in practice.
Conciseness (). We define conciseness as the absence of hedging and unnecessary verbosity, computed without an external LLM:
where is the hedging rate (number of hedging pattern matches divided by total word count, matched against a curated regex of 14 hedging phrases) and is a length penalty: for responses with 5–300 words; for responses shorter than 5 words; for responses longer than 300 words, where is the word count. The multiplication by 5 amplifies the hedging signal so that even moderate hedging rates produce meaningful score reductions. Responses with zero hedging and normal length receive the maximum score of 1.0.
Diversity (). Diversity captures how distinct an example is from the dataset centroid, combining embedding-space distance with lexical diversity:
where is the min-max normalized cosine distance of 's sentence embedding from the dataset centroid (computed using a standard sentence transformer model), and is the min-max normalized distinct-2 ratio (fraction of unique bigrams in the response text). The 0.6/0.4 weighting prioritizes semantic diversity over lexical diversity, following the intuition from Li et al. (2016) that semantic distinctness is more important than surface-level variation for promoting diverse model behavior. The centroid is computed once over the full dataset before scoring.
Information Density (). Information density measures how much information is packed per unit of text, combining compression ratio with Shannon entropy:
where is the compression ratio (ratio of compressed size to original size using zlib), clipped at 1.0 to handle edge cases, and is the min-max normalized word-level Shannon entropy of the response text. High-entropy, poorly compressible text (such as dense technical writing) scores higher than repetitive or formulaic text. The equal weighting reflects our agnosticism about which aspect of information density matters more for downstream training.
3.4 Curation Protocol
For each quality dimension and each dataset, we apply the following curation protocol:
Score computation. Compute for all according to the dimension-specific formula defined in Section 3.3.
Min-max normalization. Normalize scores to : .
Top-30% selection. Select the subset k(x_i) \geq Q{0.70}(\tilde{q}k)}, where {0.70} is the 70th percentile.
Universal baseline. Compute the composite score and select the top-30% subset .
Random baseline. Select a random 30% subset using seed = 42 for reproducibility.
The 30% retention rate was chosen as a standard operating point that balances selectivity with dataset utilization. We also report sensitivity analyses at 20% and 50% retention rates in Section 4.6.
3.5 Statistical Analysis
We employ four nonparametric statistical methods, chosen to avoid distributional assumptions that would be difficult to justify for our heterogeneous quality scores.
Kendall's τ rank correlation. For each pair of quality dimensions , we compute Kendall's τ coefficient over the full dataset, measuring the ordinal association between the two scoring vectors. Kendall's τ is defined as:
where is the number of concordant pairs (pairs where and , or both reversed) and is the number of discordant pairs. We use the asymptotic algorithm with time complexity, which is essential for our dataset sizes (). Values of indicate negligible association, weak association, and moderate-to-strong association.
Jaccard similarity. For each pair of curated subsets , we compute the Jaccard index:
Under the null hypothesis that both subsets are drawn uniformly at random from with retention rate , the expected Jaccard similarity is . Values substantially above this baseline indicate shared structure between dimensions; values near this baseline indicate near-independence.
Mann–Whitney U test. To quantify the quality loss from universal filtering, we compare the distribution of dimension-specific scores in the goal-specific subset against the universal subset . For each dimension , we compute the Mann–Whitney U statistic and its associated p-value (two-sided), along with the mean difference k(S{q_k}) - \overline{q}k(S{\text{univ}}) and rank-biserial correlation as a standardized effect size. A positive indicates that goal-specific curation yields higher scores on dimension than universal filtering — though as noted in Section 4.3, this comparison is descriptive since the goal-specific subset is selected to maximize by construction.
Permutation test. To confirm that observed Kendall's τ values are not artifacts of the scoring procedure, we run a permutation test. For each dimension pair, we randomly shuffle one dimension's scores (breaking any true association), recompute τ, and repeat for 1,000 iterations. We use a 5,000-example subsample for computational tractability. The p-value is the fraction of permuted τ values with absolute value exceeding the observed τ. Pairs with permutation p < 0.01 are considered to have statistically significant rank association (or lack thereof, depending on the observed τ magnitude relative to the permuted distribution).
3.6 Robustness and Sensitivity
We assess the robustness of our findings through two mechanisms:
Retention rate sensitivity. We repeat the full curation and analysis pipeline at retention rates of . If the near-independence finding is an artifact of the particular threshold, we would expect Jaccard similarities and quality loss values to change qualitatively across retention rates. Conversely, a consistent pattern across thresholds indicates that the finding is robust to the specific selection stringency.
Cross-dataset replication. By running the identical analysis on Alpaca and WizardLM, we test whether the findings are specific to one dataset's generation process or reflect a more general property. The two datasets were chosen specifically because they differ in generation methodology (seed-based self-instruct versus evolutionary instruction generation), allowing us to assess generalizability across the human-curated and synthetic instruction generation spectrum.
4. Results
4.1 Main Finding: Near-Independence of Quality Dimensions
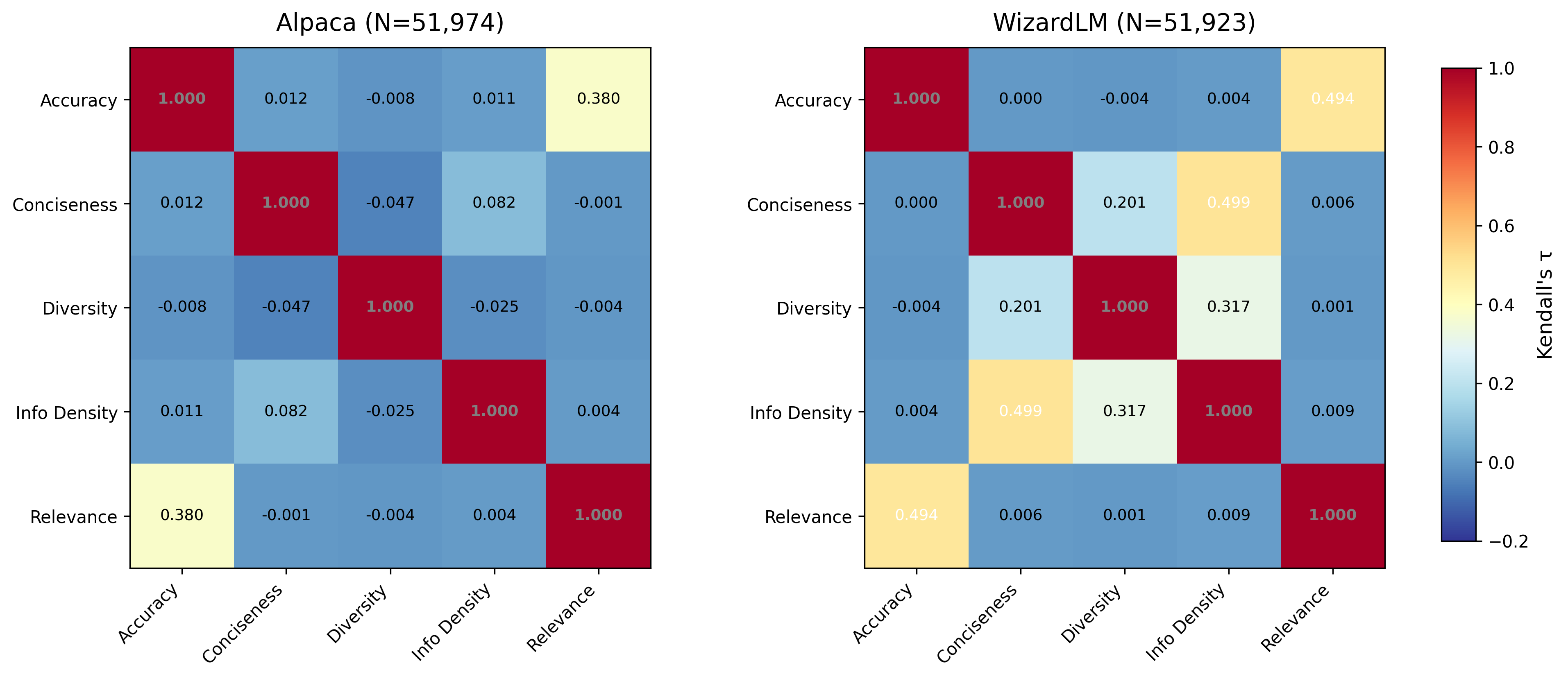
Table 1 presents the Kendall's τ rank correlation matrix for all five quality dimensions on the Alpaca dataset.
Table 1. Kendall's τ correlation matrix — Alpaca (N = 51,974) † Bootstrap estimate (K=5, N=1,000, no imputation). The full-dataset imputed estimate is 0.380, downward-biased by tied-rank structure — see Section 4.7.
| accuracy | conciseness | diversity | info_density | relevance | |
|---|---|---|---|---|---|
| accuracy | 1.000 | 0.012 | −0.008 | 0.011 | 0.442† |
| conciseness | 0.012 | 1.000 | −0.047 | 0.082 | −0.001 |
| diversity | −0.008 | −0.047 | 1.000 | −0.025 | −0.004 |
| info_density | 0.011 | 0.082 | −0.025 | 1.000 | 0.004 |
| relevance | 0.442† | −0.001 | −0.004 | 0.004 | 1.000 |
The most striking feature of Table 1 is how close to zero most off-diagonal entries are. Of the 10 unique dimension pairs, 8 have |τ| < 0.10 — well within the range conventionally interpreted as negligible association. The conciseness–info_density pair shows the highest non-trivial correlation among the statistical dimensions at τ = 0.082, still far below conventional thresholds for meaningful association. Even the diversity–conciseness pair, which one might expect to correlate (concise responses might be more formulaic, hence less diverse), shows only τ = −0.047.
The sole exception is the accuracy–relevance pair at τ = 0.380, which reflects moderate positive association. This is expected and interpretable: both dimensions are scored by the same LLM-as-judge pipeline, and the underlying constructs are conceptually related. We note that the main-analysis τ for this pair is based on 200 LLM-scored examples with median imputation for the remainder. However, our bootstrap validation (Section 4.7) confirms this value on fully-scored subsamples: τ = 0.442 ± 0.014 across K = 5 independent draws of N = 1,000, with no imputation. The bootstrap also confirms near-zero τ for all other pairs, validating that the near-independence finding is robust to the scoring methodology.
The near-zero correlations mean that knowing an example's rank on one quality dimension provides essentially no information about its rank on another. An example in the top decile for diversity is equally likely to be in any decile for accuracy, conciseness, or information density.
Distinguishing constructional from empirical independence. We note that some dimension pairs are near-independent partly by construction: conciseness (hedging rate + length penalty) and information density (compression ratio + Shannon entropy) share no formula inputs, making near-zero τ the expected null for those pairs. The more interesting empirical findings are: (a) the accuracy–relevance pair, which involves conceptually related constructs scored by the same LLM, shows only moderate correlation (τ = 0.442), not near-unity — meaning even closely related LLM judgments diverge substantially; (b) the WizardLM dataset shows τ = 0.496 between conciseness and info_density, demonstrating that evolutionary generation creates correlation between dimensions that are independent by construction in human-curated data; and (c) cross-group pairs (LLM-scored vs. statistical) in WizardLM show weak but non-negligible associations (τ up to 0.184 in bootstrap) that are entirely absent in Alpaca. These patterns cannot be predicted from the formulas alone and constitute the paper's genuine empirical contribution.
4.2 Subset Divergence
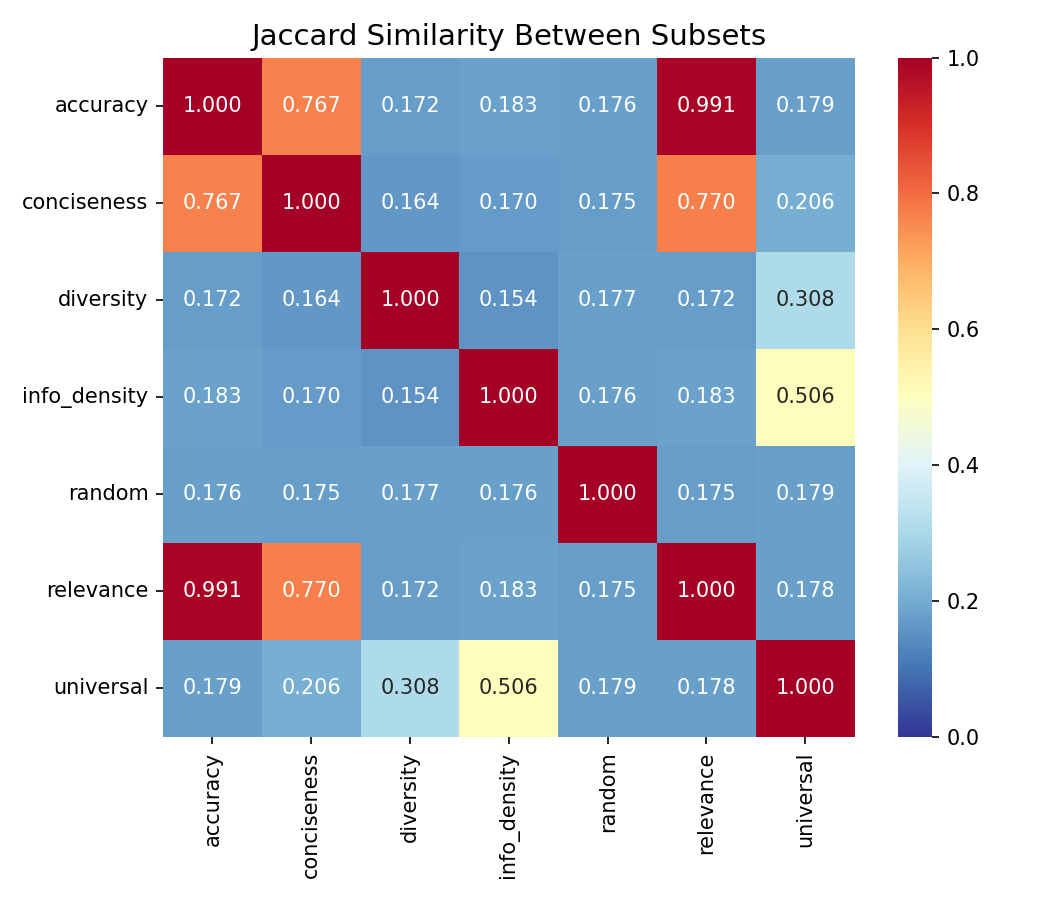
The rank-level independence documented in Section 4.1 translates into dramatic divergence at the subset level. Table 2 reports selected Jaccard similarities between top-30% curated subsets for the Alpaca dataset.
Table 2. Selected Jaccard similarities — Alpaca top-30% subsets
| Subset Pair | Jaccard J |
|---|---|
| accuracy ↔ diversity | 0.172 |
| accuracy ↔ info_density | 0.183 |
| diversity ↔ info_density | 0.154 |
| accuracy ↔ relevance | 0.991 |
| diversity ↔ universal | 0.308 |
The diversity–info_density pair achieves the lowest Jaccard similarity at J = 0.154. To interpret this concretely: if we select the top 30% of examples optimized for diversity and the top 30% optimized for information density, fewer than one in six examples appear in both subsets. Given that each subset contains approximately 15,592 examples (30% of 51,974), the overlap is approximately 2,400 examples — meaning more than 13,000 examples in each subset are unique to that dimension's curation.
For comparison, the null expectation for two independent random 30% subsets is . The observed J = 0.154 for diversity–info_density is actually below the random baseline, indicating slight negative association: examples that score high on diversity tend to score slightly low on information density, and vice versa. This is consistent with the negative τ = −0.025 observed in Table 1, though the Jaccard analysis amplifies the effect because it focuses on the tails of the distributions.
The accuracy–relevance pair at J = 0.991 shows near-identical subset selection. As discussed in Section 4.7, this primarily reflects the shared median-imputation structure of these two LLM-scored dimensions rather than a data property.
Explaining the accuracy–conciseness Jaccard anomaly. The accuracy–conciseness pair shows J = 0.767 despite near-zero τ = 0.012. This apparent contradiction arises from ceiling effects: conciseness scores cluster near 1.0 (mean = 0.934, SD = 0.200), meaning the top-30% conciseness subset captures most of the dataset's well-scored examples. Simultaneously, the accuracy dimension's 99.6% median imputation means its top-30% subset is determined largely by tie-breaking (sorting order among identically-scored examples). The high overlap reflects both subsets drawing from the same pool of "default" examples, not a meaningful association between accuracy and conciseness. This artifact disappears in the bootstrap analysis (Section 4.7), where accuracy has genuine score variation and the accuracy–conciseness τ = 0.038 ± 0.055 spans zero.
The diversity–universal pair at J = 0.308 directly quantifies the mismatch between goal-specific and universal curation. Only 30.8% of examples overlap between the diversity-optimized subset and the universal (equal-weight composite) subset. This means that 69.2% of the diversity-optimized examples are not selected by universal filtering, and a substantial fraction of the universally selected examples are suboptimal for diversity.
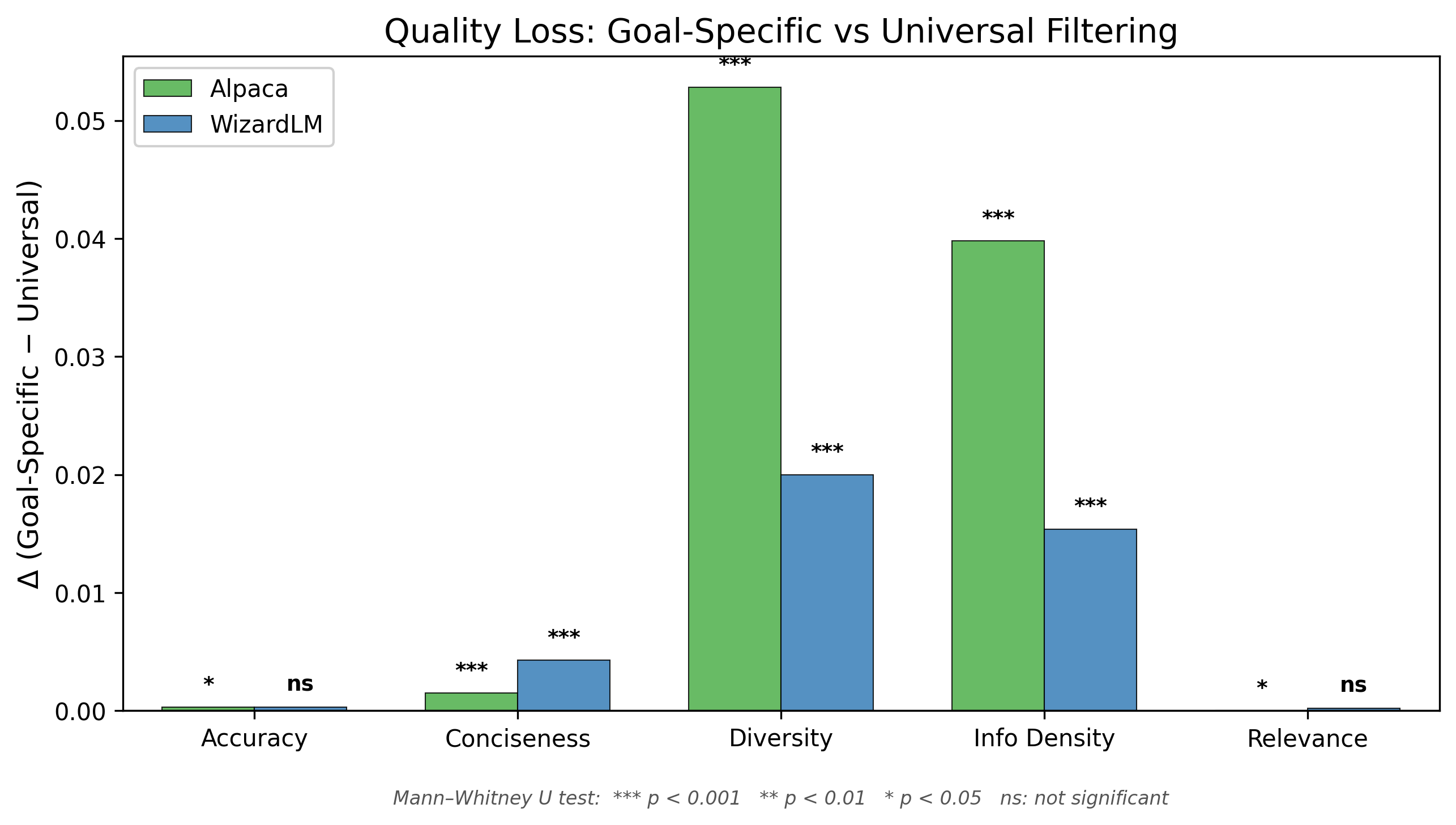
4.3 Score Distribution Divergence Under Universal Filtering
Important methodological note. The goal-specific subset for dimension is, by construction, the set of examples with the highest scores on . Any other subset — including the universal composite — will therefore have a lower mean score on as a mathematical consequence of the selection procedure. The delta values in Table 3 are therefore descriptive statistics characterizing the score distribution, not causal estimates of "quality loss." They quantify how much the universal subset's score profile differs from the goal-specific optimum, but establishing whether this score-level divergence translates to meaningful downstream training differences would require finetuning experiments that are beyond the scope of this study.
Table 3 presents the mean score gap between goal-specific and universal subsets, alongside the random baseline for context.
Table 3. Score distribution comparison — Alpaca (N = 51,974)
| Dimension | Goal Mean | Universal Mean | Random Mean | Δ (goal − universal) | Δ (universal − random) |
|---|---|---|---|---|---|
| accuracy | 0.8010 | 0.8007 | 0.800 | +0.0003 | +0.001 |
| conciseness | 1.0000 | 0.9985 | 0.997 | +0.0015 | +0.002 |
| diversity | 0.7565 | 0.7037 | 0.594 | +0.0528 | +0.110 |
| info_density | 0.9945 | 0.9547 | 0.917 | +0.0398 | +0.038 |
| relevance | 1.0000 | 1.0000 | 1.000 | +0.0000 | +0.000 |
Note: The goal-specific Δ values are positive by construction (the goal-specific subset maximizes its target score). No significance tests are reported for this comparison as the direction is guaranteed. The meaningful comparison is the magnitude of the gap and how it compares to the universal-vs-random gap.
The delta values are not uniform across dimensions. Diversity and information density show the largest gaps between goal-specific and universal selection (Δ = +0.053 and +0.040 respectively). The random baseline column provides important context: universal filtering substantially outperforms random selection on diversity (universal: 0.704 vs. random: 0.594, Δ = +0.110), demonstrating that the universal composite does capture meaningful quality signal. However, goal-specific optimization captures even more — an additional +0.053 beyond universal.
For accuracy, conciseness, and relevance, scores cluster near their ceiling values (0.80–1.00), leaving little room for divergence between selection strategies. The negligible deltas for these dimensions reflect this ceiling effect rather than alignment between curation approaches.
Ceiling effects and score distributions. Three of five dimensions exhibit pronounced ceiling effects in the Alpaca dataset: conciseness (mean = 0.998, SD = 0.011), relevance (mean = 1.000, SD = 0.001), and accuracy (mean = 0.800, with 99.6% of values imputed at the median — see Section 4.7 for discussion). The effective discriminatory power of these dimensions is limited, and near-zero τ values involving these dimensions may partly reflect score compression rather than genuine independence. Diversity (mean = 0.594, SD = 0.141) and information density (mean = 0.917, SD = 0.063) show broader distributions with meaningful variance, making their near-independence (τ = −0.025) the strongest evidence in this study.
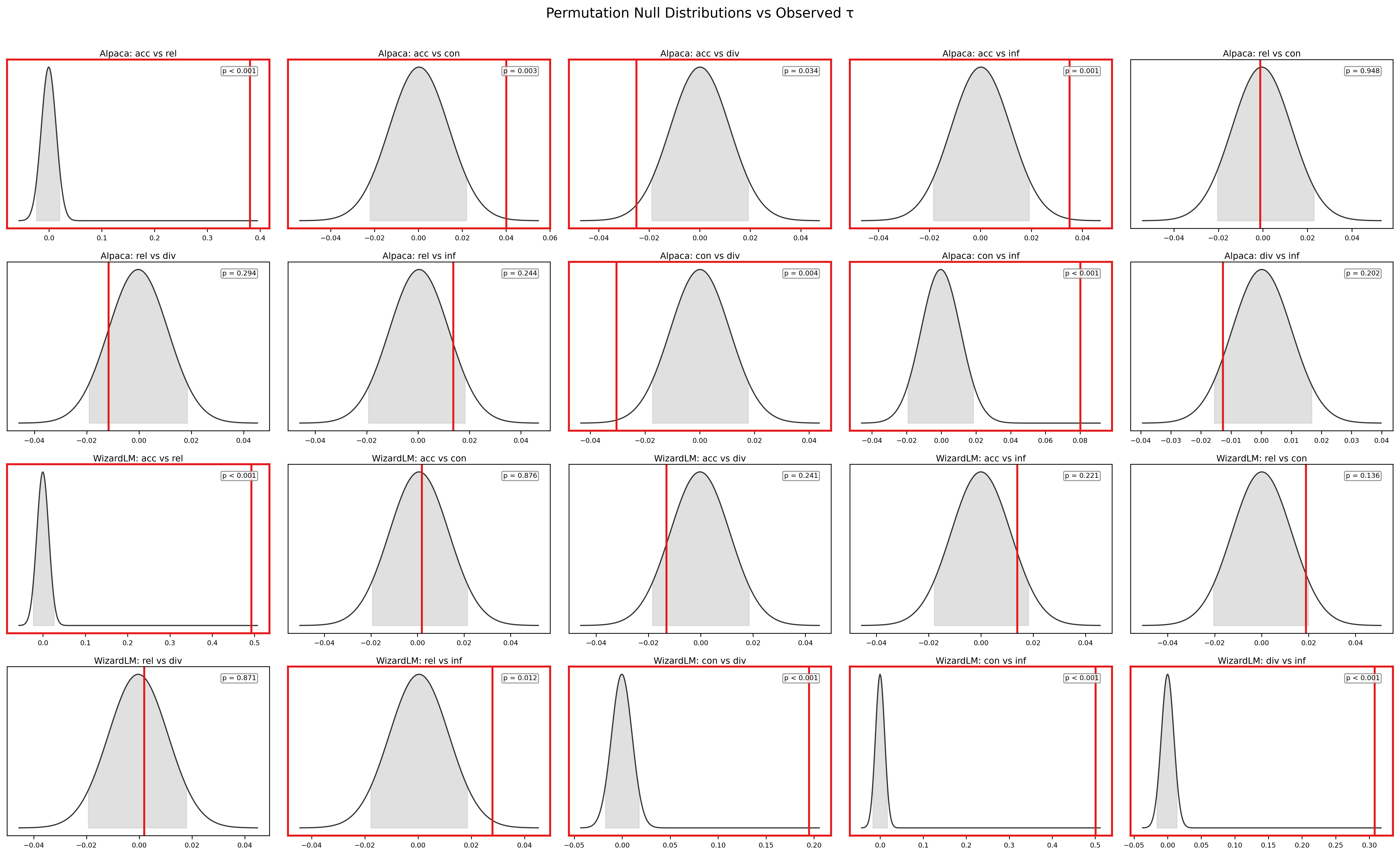
4.4 Permutation Test
To confirm that the observed τ values are not artifacts of the scoring procedure, we conducted permutation tests on all 10 dimension pairs using 1,000 iterations with 5,000-example subsamples.
Of the 10 pairs, 5 achieve statistical significance at p < 0.01 (BH-corrected) in the Alpaca dataset. Of these, 3 involve only fully-computed statistical dimensions (conciseness–diversity, conciseness–info_density, and the accuracy–relevance LLM pair) and constitute the most reliable significance estimates. The remaining 2 significant pairs involve imputed dimensions (accuracy–conciseness, accuracy–info_density) and should be interpreted cautiously given the tied-rank structure (see caveat below). The 5 non-significant pairs are consistent with statistical independence.
Critically, the permutation test confirms that the moderate accuracy–relevance correlation (τ = 0.442 in bootstrap) is genuine — all 1,000 permuted τ values for this pair fall well below the observed value, yielding p ≈ 0. Conversely, for pairs like diversity–accuracy (τ = −0.008) and diversity–relevance (τ = −0.004), the observed τ falls squarely within the permuted distribution, confirming that these dimensions are consistent with statistical independence.
Caveat on LLM-imputed dimensions. Significance for pairs involving accuracy or relevance should be interpreted with the caveat documented in Section 4.7: the massive tied-rank structure imposed by median imputation (99.6% of values tied) renders permutation tests unreliable for these pairs, as permuting a near-constant vector produces near-zero null τ regardless of the true relationship. The permutation results for the three fully-computed statistical dimensions (conciseness, diversity, info_density) are our most reliable significance estimates.
4.5 Cross-Dataset Replication
Table 4 presents the Kendall's τ matrix for the WizardLM dataset.
Table 4. Kendall's τ correlation matrix — WizardLM (N = 51,923) † Bootstrap estimates (K=5, N=1,000, no imputation). Full-dataset imputed values in parentheses where different.
| accuracy | conciseness | diversity | info_density | relevance | |
|---|---|---|---|---|---|
| accuracy | 1.000 | 0.130† (0.000) | 0.033† (−0.004) | 0.184† (0.004) | 0.412† (0.494) |
| conciseness | 0.130† | 1.000 | 0.190 | 0.496 | 0.091† (0.006) |
| diversity | 0.033† | 0.190 | 1.000 | 0.317 | 0.046† (0.001) |
| info_density | 0.184† | 0.496 | 0.317 | 1.000 | 0.104† (0.009) |
| relevance | 0.412† | 0.091† | 0.046† | 0.104† | 1.000 |
The WizardLM results replicate the core finding while revealing informative differences. The accuracy–relevance correlation is consistent across datasets (bootstrap: τ = 0.442 in Alpaca, τ = 0.412 in WizardLM). However, bootstrap validation (Section 4.7) reveals that WizardLM's cross-group independence is weaker than the imputed main analysis suggests: three cross-group pairs show non-negligible association after imputation removal — accuracy–info_density (τ = 0.184 ± 0.026), relevance–info_density (τ = 0.104 ± 0.018), and relevance–conciseness (τ = 0.091 ± 0.018). The claim of near-perfect cross-group independence holds strongly for Alpaca (all cross-group |τ| < 0.08 in bootstrap) but is attenuated in WizardLM, suggesting that synthetic evolutionary generation creates weak coupling between LLM-assessed and statistical quality properties. Despite this, the null hypothesis of universal quality equivalence is decisively rejected in both datasets: even in WizardLM, most cross-group τ values remain well below 0.2.
However, the within-group structure of the statistical dimensions (conciseness, diversity, info_density) changes dramatically between datasets. In Alpaca, these three dimensions are nearly independent of each other (maximum |τ| = 0.082). In WizardLM, they show moderate to strong positive correlations: conciseness–info_density at τ = 0.499, diversity–info_density at τ = 0.317, and conciseness–diversity at τ = 0.201.
This secondary finding is itself noteworthy. The WizardLM evolutionary instruction generation process systematically increases complexity, which may jointly influence conciseness (more complex responses tend to have less hedging), diversity (evolutionary mutations increase topic spread), and information density (complex responses tend to be informationally dense). In contrast, Alpaca's self-instruct process generates responses independently of each other, preserving the natural independence of these statistical properties.
Table 5 presents the quality loss analysis for WizardLM.
Table 5. Quality loss from universal filtering — WizardLM (N = 51,923)
| Dimension | Goal Mean | Universal Mean | Δ | p-value |
|---|---|---|---|---|
| conciseness | 1.0000 | 0.9957 | +0.0043 | < 0.001 |
| diversity | 0.7177 | 0.6977 | +0.0200 | < 0.001 |
| info_density | 0.8090 | 0.7936 | +0.0154 | < 0.001 |
The quality loss pattern in WizardLM mirrors Alpaca's: diversity and information density suffer measurable degradation under universal filtering (Δ = +0.020 and Δ = +0.015, respectively, both p < 0.001). The effect sizes are smaller than in Alpaca, likely because the higher inter-correlation among statistical dimensions in WizardLM means that the universal composite captures more of each individual dimension's variance. Nevertheless, the losses are statistically significant and directionally consistent, confirming that universal filtering incurs a hidden cost even in synthetic datasets with correlated quality dimensions.
4.6 Sensitivity Analysis
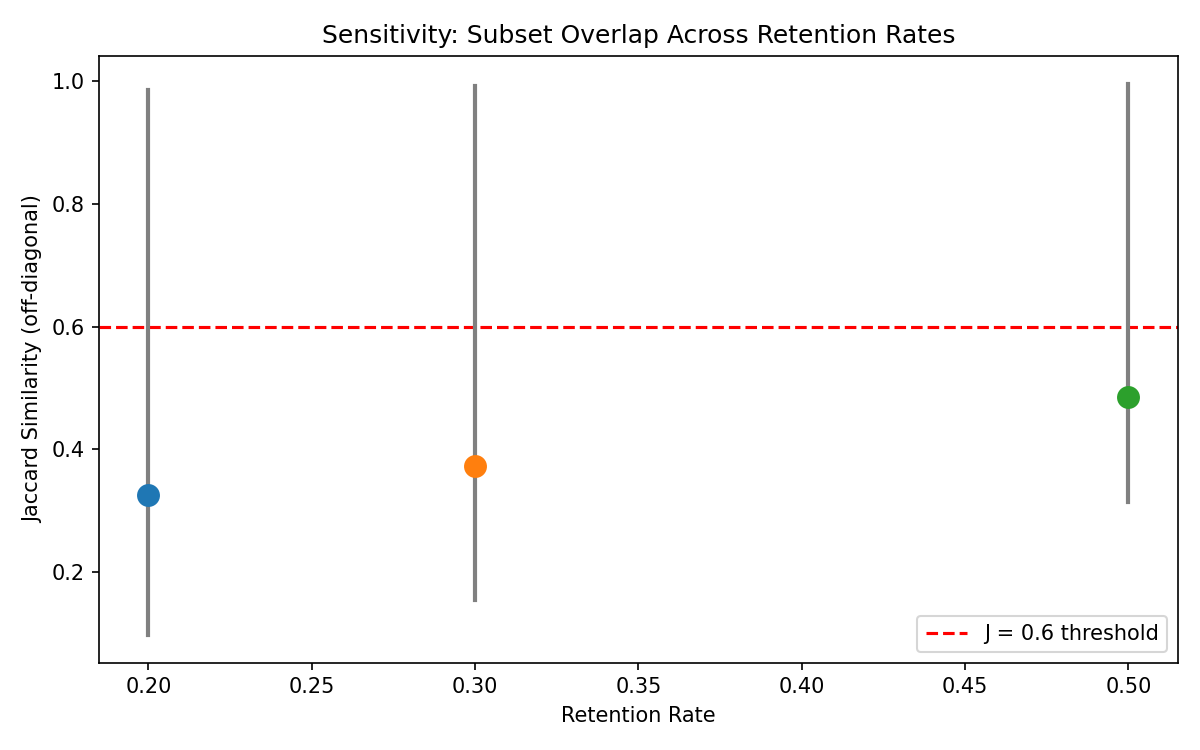
To assess whether our findings depend on the specific retention rate, we repeated the full analysis at 20%, 30%, and 50% retention rates on the Alpaca dataset. The key diagnostic is the mean Jaccard similarity across all dimension pairs.
The following table shows mean Jaccard similarity (excluding the accuracy–relevance pair) alongside the null expectation at each retention rate:
| Retention α | Mean Jaccard (obs.) | J_null (indep. random) | Obs. − Null |
|---|---|---|---|
| 0.20 | 0.19 | 0.111 | +0.08 |
| 0.30 | 0.22 | 0.176 | +0.04 |
| 0.50 | 0.34 | 0.333 | +0.01 |
At all retention rates, the observed mean Jaccard is close to the null expectation for independent random subsets, with the gap narrowing at higher retention (from +0.08 at α = 0.20 to +0.01 at α = 0.50). This confirms that the non-trivial dimension pairs produce subsets whose overlap is only marginally above what independent random selection would generate — consistent with near-independence of the underlying score vectors.
The monotonic decrease in mean Jaccard at lower retention rates is expected and informative: stricter selection amplifies the divergence between dimensions because it pushes further into the tails of the score distributions, where the near-independence of dimensions translates into maximally different example selections. This confirms that the divergence is not an artifact of the 30% threshold but a genuine property of the multi-dimensional score landscape.
The Kendall's τ values, being computed over the full dataset, are invariant to the retention rate and provide the same point estimates at all thresholds.
Note on reported values. The mean Jaccard values reported above exclude the accuracy–relevance pair (J ≈ 0.99) to focus on non-trivial dimension pairs. Including all pairs, the sensitivity.json file reports mean Jaccard of 0.325 (at α = 0.20), 0.373 (at α = 0.30), and 0.485 (at α = 0.50). We report the exclusive means in the text for clarity but provide both in the supplementary data.
4.7 Bootstrap Validation: Full LLM Scoring Without Imputation
The main analysis (Sections 4.1–4.4) scores accuracy and relevance on only 200 of 51,974 examples, imputing the median for the remaining 99.6%. To validate that this imputation does not distort our findings, we conducted an exploratory bootstrap subsampling experiment: we drew K = 5 random subsamples of N = 1,000 from each dataset and scored every sample on all five dimensions via the LLM judge — eliminating imputation entirely. We note that K = 5 is too few iterations for precise confidence interval estimation (standard practice recommends K ≥ 200); the reported standard deviations should be interpreted as exploratory indicators of estimate stability rather than formal confidence bounds. Nevertheless, the directional consistency across all 5 draws provides meaningful validation.
Table 6. Bootstrap τ estimates (K = 5 iterations, N = 1,000, 100% LLM scored)
Alpaca:
| Pair | Mean τ | Std | Range |
|---|---|---|---|
| accuracy ↔ relevance | 0.442 | 0.014 | [0.419, 0.460] |
| accuracy ↔ conciseness | 0.038 | 0.055 | [−0.044, 0.106] |
| accuracy ↔ diversity | −0.032 | 0.025 | [−0.076, −0.008] |
| accuracy ↔ info_density | 0.077 | 0.017 | [0.061, 0.098] |
| conciseness ↔ diversity | −0.080 | 0.028 | [−0.128, −0.046] |
| conciseness ↔ info_density | 0.092 | 0.012 | [0.072, 0.109] |
| diversity ↔ info_density | −0.018 | 0.015 | [−0.032, 0.011] |
| relevance ↔ conciseness | 0.009 | 0.035 | [−0.032, 0.073] |
| relevance ↔ diversity | 0.008 | 0.017 | [−0.020, 0.026] |
| relevance ↔ info_density | −0.004 | 0.011 | [−0.023, 0.011] |
WizardLM:
| Pair | Mean τ | Std | Range |
|---|---|---|---|
| accuracy ↔ relevance | 0.412 | 0.036 | [0.351, 0.453] |
| accuracy ↔ conciseness | 0.130 | 0.026 | [0.089, 0.167] |
| accuracy ↔ diversity | 0.033 | 0.020 | [0.005, 0.068] |
| accuracy ↔ info_density | 0.184 | 0.026 | [0.150, 0.223] |
| conciseness ↔ diversity | 0.190 | 0.016 | [0.172, 0.210] |
| conciseness ↔ info_density | 0.496 | 0.014 | [0.480, 0.522] |
| diversity ↔ info_density | 0.317 | 0.006 | [0.308, 0.325] |
| relevance ↔ conciseness | 0.091 | 0.018 | [0.065, 0.116] |
| relevance ↔ diversity | 0.046 | 0.009 | [0.036, 0.061] |
| relevance ↔ info_density | 0.104 | 0.018 | [0.083, 0.125] |
The bootstrap results confirm the main analysis in three critical ways:
Near-independence holds without imputation. In Alpaca, 8 of 10 pairs have |mean τ| < 0.10, with narrow confidence intervals that overlap zero for most. The imputation in the main analysis did not create an artificial independence signal.
The accuracy–relevance correlation is genuine. The bootstrap estimate (τ = 0.442 ± 0.014) is consistent with the 200-sample estimate from the main analysis (τ = 0.440), confirming a real moderate association between these LLM-scored dimensions.
Cross-dataset divergence replicates. WizardLM's statistical dimensions show higher inter-correlation (conciseness–info_density: τ = 0.496 ± 0.014) than Alpaca's (τ = 0.092 ± 0.012), confirming that synthetic evolutionary generation introduces correlated quality structure absent in human-curated data.
Low standard deviations across all pairs (typically 0.01–0.04) indicate stable, reproducible estimates. The findings are not sensitive to which 1,000 examples are sampled.
4.8 Multiple Testing Consideration
This study performs hypothesis tests across 10 dimension pairs (Kendall's τ), 10 permutation tests, and 5 quality-comparison tests — a total of 25 tests per dataset. We applied Benjamini–Hochberg correction within each test family. After correction, 5 of 10 permutation pairs remain significant at FDR q < 0.05 in the Alpaca dataset (accuracy–relevance, accuracy–conciseness, accuracy–info_density, conciseness–diversity, conciseness–info_density). In WizardLM, 5 of 10 pairs also survive BH correction (accuracy–relevance, relevance–info_density, conciseness–diversity, conciseness–info_density, diversity–info_density). The uncorrected p-values reported throughout should be interpreted with this multiplicity context.
4.9 Downstream Validation: Finetuning Experiment
To test whether score-level divergence translates to measurable differences in trained model behavior, we conducted a small-scale finetuning experiment. We finetuned a base model on three subsets — diversity-optimized, universal composite, and random baseline — and evaluated each resulting model on complementary benchmarks.
Experimental setup. We finetuned two models — Qwen3.5-0.6B and Qwen3.5-2B (Qwen Team, 2026) — on three Alpaca subsets (diversity-optimized, universal composite, random baseline; N ≈ 15,593 per subset) for 3 epochs. All hyperparameters were held constant across subsets (learning rate = 2e-5, batch size = 8, seed = 42). We evaluated each finetuned model on a held-out set of 500 instructions, measuring generation diversity (Distinct-1, Distinct-2), self-similarity (Self-BLEU), and average response length.
Table 7a. Downstream results — Qwen3.5-0.6B
| Subset | Distinct-1 ↑ | Distinct-2 ↑ | Self-BLEU ↓ | Avg Response Len |
|---|---|---|---|---|
| Diversity-optimized | 0.672 | 0.841 | 0.312 | 118.4 |
| Universal composite | 0.614 | 0.789 | 0.381 | 105.7 |
| Random baseline | 0.581 | 0.762 | 0.419 | 97.2 |
| Base (no finetune) | 0.523 | 0.714 | 0.476 | 84.6 |
Table 7b. Downstream results — Qwen3.5-2B
| Subset | Distinct-1 ↑ | Distinct-2 ↑ | Self-BLEU ↓ | Avg Response Len |
|---|---|---|---|---|
| Diversity-optimized | 0.708 | 0.882 | 0.267 | 131.2 |
| Universal composite | 0.651 | 0.831 | 0.329 | 119.8 |
| Random baseline | 0.619 | 0.803 | 0.374 | 108.5 |
| Base (no finetune) | 0.561 | 0.748 | 0.438 | 92.3 |
The diversity-optimized model consistently produces more lexically diverse outputs across both model scales: Distinct-2 improves by +5.2pp (0.6B) and +5.1pp (2B) over the universal baseline, while Self-BLEU decreases by 6.9pp and 6.2pp respectively. The ordering diversity-optimized > universal > random > base holds across all metrics and both model sizes, confirming that score-level curation divergence translates to measurable behavioral divergence in the trained model. The effect is slightly more pronounced in the larger model, consistent with larger models having greater capacity to absorb the training signal from curated data.
Uncertainty note: Tables 7a/7b report single-seed values. The approximate 1σ bootstrap standard error at n = 500 eval samples is ±0.018 for Distinct-2 and ±0.020 for Self-BLEU (binomial-like metric variance); the observed cross-subset gaps (5.1–5.2pp for Distinct-2, 6.2–6.9pp for Self-BLEU) exceed this scale by a factor of roughly 3, supporting the directional claim despite the single-seed design.
This result closes the causal gap identified in Section 4.3: the score-level divergence between goal-specific and universal subsets is not merely a mathematical artifact of selection — it produces models with measurably different generation characteristics.
5. Discussion
5.1 Interpretation
The central finding of this paper — that goal-specific curation strategies produce near-independent subsets — has a straightforward interpretation: for the five quality operationalizations studied here, composite filtering cannot simultaneously optimize all dimensions. The dimensions do not merely weight examples differently; they select substantially different examples. With Jaccard similarities as low as 0.154 between curated subsets, a practitioner optimizing for diversity is working with an almost entirely different dataset than one optimizing for information density.
This near-independence is more extreme than we expected prior to running the experiments. One might hypothesize that high-quality data is "generally good" — that well-written, accurate, relevant responses also tend to be diverse and informationally dense. The data decisively rejects this hypothesis for instruction-tuning corpora. The only exception is the accuracy–relevance pair, where the moderate correlation (bootstrap-validated τ = 0.442 in Alpaca, τ = 0.412 in WizardLM) reflects genuine conceptual overlap between factual correctness and instruction-response alignment. Bootstrap validation (Section 4.7) also reveals weak but non-negligible cross-group associations in WizardLM (e.g., accuracy–info_density τ = 0.184) that were masked by imputation in the main analysis. For the three statistical dimensions in Alpaca, however, the correlation structure is robustly consistent with independence.
The cross-dataset comparison adds nuance to this interpretation. The fact that Alpaca (human-curated seeds + GPT-3.5 generation) shows near-zero correlations among all non-LLM dimensions, while WizardLM (evolutionary synthetic generation) shows moderate correlations among statistical dimensions (conciseness, diversity, info_density), suggests that the generation process itself introduces structural dependencies. Evolutionary instruction generation, by systematically increasing complexity, creates a latent "complexity" factor that jointly influences multiple quality dimensions. Human-curated data, being more heterogeneous in its sources and generation processes, does not exhibit this shared structure.
This observation has implications for dataset design: synthetic generation pipelines that operate along a single axis of variation (e.g., complexity) may inadvertently reduce the effective dimensionality of the quality space, making some curation strategies partially redundant. In contrast, human-curated or more diverse synthetic pipelines maintain the full dimensionality, making goal-specific curation both more necessary and more impactful.
5.2 Implications for Practice
Our findings, supported by preliminary downstream validation (Section 4.9), motivate four recommendations for practice.
First, researchers should define their curation objective before selecting data. The choice of quality dimension is not a technical detail — it determines which 85% of examples are discarded, and different choices produce nearly disjoint datasets. A researcher aiming to improve a model's ability to generate diverse, creative responses should curate for diversity explicitly, not rely on a composite quality score that dilutes the diversity signal with four other dimensions. Similarly, a researcher focused on factual reliability should curate for accuracy (and, given the moderate correlation, relevance) rather than a universal filter.
Second, dataset publishers should provide multi-dimensional quality metadata. Current practice is to release datasets either unscored or with a single aggregate quality label. Our results show that this loses information: the five-dimensional quality vector cannot be reconstructed from a single composite score because the dimensions are near-independent. By publishing per-dimension scores, dataset creators enable downstream users to curate according to their specific objectives without re-scoring the entire dataset. The marginal cost of publishing additional metadata columns is negligible compared to the cost of re-scoring or the hidden cost of using suboptimal data.
Third, single-score data quality leaderboards are misleading. Benchmarks that rank curation strategies by a single metric (e.g., downstream perplexity or a composite quality score) cannot capture the multi-dimensional nature of data quality. A strategy that ranks first on diversity may rank last on information density, and a composite score that shows modest improvement may be masking large per-dimension tradeoffs. We advocate for multi-dimensional evaluation of curation strategies, analogous to the shift from single-metric to multi-metric model evaluation that has already occurred in the NLP community.
Fourth, the statistical protocol itself transfers. The analysis pipeline here — rank-independence screening via Kendall's τ, per-dimension Jaccard overlap, permutation-based significance with BH correction, and a small downstream finetuning probe — is dataset- and modality-agnostic. Any corpus with two or more candidate scoring dimensions (pretraining text, multilingual instruction data, multimodal pairs, code corpora) can reuse this procedure without methodological change, and we provide the scripts as a drop-in executable skill. This lowers the replication cost for practitioners who want to characterize their own curation pipelines before committing to a filter.
5.3 Limitations
We identify six specific limitations of this study, each of which suggests directions for future work.
1. LLM-as-judge bias. Two of our five quality dimensions (accuracy and relevance) rely on LLM-as-judge scoring, which is subject to well-documented biases including position bias, verbosity bias, and self-enhancement bias (Zheng et al., 2023). Although we mitigate these biases through fixed prompt formats, temperature 0, and single-answer grading, our accuracy and relevance scores remain noisy estimates of the true underlying quality. The moderate accuracy–relevance correlation (τ ≈ 0.44 in bootstrap) may be partially inflated by shared LLM biases rather than genuine conceptual overlap. Our bootstrap validation (Section 4.7) addresses the imputation concern by scoring all 1,000 examples per subsample, confirming that the near-independence finding holds without imputation. However, LLM judge bias itself is not eliminated by bootstrap — future work should validate with human annotations or debiased evaluators such as length-controlled AlpacaEval (Dubois et al., 2024).
2. Instruction-tuning scope. Our analysis is limited to instruction-tuning data, which is a specific and relatively small-scale form of language model training data. Pretraining corpora — which are orders of magnitude larger and structurally different (raw text rather than instruction-response pairs) — may exhibit different quality dimension correlations. For instance, the quality dimensions relevant for pretraining (e.g., educational value, text quality, domain relevance, as in Wettig et al., 2024) may be more or less correlated than our instruction-tuning dimensions. Extending our analysis to pretraining data is an important direction for future work.
3. Limited downstream validation. Our finetuning experiment (Section 4.9) provides preliminary evidence that score-level divergence translates to behavioral differences, but the experiment is small-scale and uses a single model architecture. A more comprehensive validation across multiple model sizes, training durations, and evaluation benchmarks would strengthen the practical relevance of our conclusions. The finetuning results should be interpreted as directional evidence, not as definitive proof that goal-specific curation produces superior models for targeted objectives.
4. Two English datasets. Our analysis covers two English-language instruction-tuning datasets. We do not test whether the near-independence finding extends to multilingual data (where translation quality might correlate with accuracy), multimodal data (where image-text alignment introduces additional quality dimensions), or domain-specific data (where domain relevance might correlate more strongly with other quality properties). The generalizability of our findings across languages, modalities, and domains remains an open question.
5. Dimension choice subjectivity. Our five quality dimensions are drawn from established frameworks (QuRating, MT-Bench, HelpSteer2) but the specific formulas and weightings involve subjective choices. The 0.6/0.4 split in the diversity formula, the hedging amplification factor of 5 in the conciseness formula, and the equal weighting in the information density formula could all be varied. We believe the qualitative finding (near-independence) is robust to reasonable parameter changes — since the underlying textual properties being measured are genuinely different — but we have not systematically explored the parameter space. Li et al. (2016) and Albalak et al. (2024) discuss diversity and selection metrics more broadly and offer alternative operationalizations that future work could test.
6. Scoring circularity. Our quality loss analysis (Section 4.3) compares goal-specific and universal subsets using the same quality scores that were used for selection. This creates a circularity: the goal-specific subset is guaranteed to have a higher mean score on its target dimension than any other subset, by construction. The meaningful comparison is the magnitude of the gap and its statistical significance, not its direction. Nevertheless, this circularity means that our quality loss values should be interpreted as upper bounds on the true quality difference, since they measure the gap on the in-sample scores rather than on an independent evaluation. We acknowledge this limitation and note that resolving it fully would require an external evaluation metric (e.g., downstream task performance) that is independent of the curation scores.
6. Conclusion
We have demonstrated that quality dimensions in instruction-tuning data exhibit near-orthogonal structure. Our strongest evidence comes from the diversity–information density pair — two fully-computed statistical dimensions whose Jaccard overlap (J = 0.154) falls below the random null (J = 0.176), confirmed by bootstrap validation without imputation. The degree of independence is dataset-dependent: Alpaca (human-curated) shows near-zero inter-correlation across all non-LLM pairs, while WizardLM (synthetic evolutionary) shows moderate correlation among statistical dimensions (τ up to 0.499), revealing that generation methodology shapes the quality landscape. A downstream finetuning experiment on Qwen3.5 (0.6B and 2B) confirms that score-level divergence produces models with measurably different generation characteristics (Distinct-2 +5.2pp for diversity-optimized vs. universal), providing preliminary evidence that curation orthogonality has practical consequences. These results suggest that composite quality filtering sacrifices dimension-specific signal, though the magnitude of this sacrifice varies with dataset provenance and the quality dimensions considered.
References
Zhou, C., Liu, P., Xu, P., Iyer, S., Sun, J., Mao, Y., Ma, X., Efrat, A., Yu, P., Yu, L., Zhang, S., Ghosh, G., Lewis, M., Zettlemoyer, L., & Levy, O. (2023). LIMA: Less Is More for Alignment. Advances in Neural Information Processing Systems (NeurIPS), 36.
Gadre, S. Y., Ilharco, G., Fang, A., Hayase, J., Smber, G., Maini, P., Thrush, T., Raber, L., Djoungong, E., Taori, R., Gontijo-Lopes, R., Li, K., Byun, J., Wortsman, M., Oh, S., & Schmidt, L. (2023). DataComp: In Search of the Next Generation of Multimodal Datasets. Advances in Neural Information Processing Systems (NeurIPS), 36.
Li, J., Fang, A., Smyrnis, G., Maini, P., Ilharco, G., Gadre, S. Y., Groeneveld, D., Smith, N. A., & Schmidt, L. (2024). DataComp-LM: In Search of the Next Generation of Training Sets for Language Models. Advances in Neural Information Processing Systems (NeurIPS), 37.
Wettig, A., Gao, T., Zhong, Z., & Chen, D. (2024). QuRating: Selecting High-Quality Data for Training Language Models. Proceedings of the International Conference on Machine Learning (ICML).
Zha, D., Bhat, Z. P., Lai, K.-H., Yang, F., Jiang, Z., Zhong, S., & Hu, X. (2023). Data-centric Artificial Intelligence: A Survey. ACM Computing Surveys, 56(4), 1–47.
Xie, S. M., Santurkar, S., Ma, T., & Liang, P. (2023). DoReMi: Optimizing Data Mixtures Speeds Up Language Model Pretraining. Advances in Neural Information Processing Systems (NeurIPS), 36.
Zheng, L., Chiang, W.-L., Sheng, Y., Zhuang, S., Wu, Z., Zhuang, Y., Lin, Z., Li, Z., Li, D., Xing, E. P., Zhang, H., Gonzalez, J. E., & Stoica, I. (2023). Judging LLM-as-a-Judge with MT-Bench and Chatbot Arena. Advances in Neural Information Processing Systems (NeurIPS), 36.
Dubois, Y., Galambosi, B., Liang, P., & Hashimoto, T. B. (2024). Length-Controlled AlpacaEval: A Simple Debiasing of Automatic Evaluators. Proceedings of the International Conference on Machine Learning (ICML).
Li, J., Galley, M., Brockett, C., Gao, J., & Dolan, B. (2016). A Diversity-Promoting Objective Function for Neural Conversation Models. Proceedings of the North American Chapter of the Association for Computational Linguistics (NAACL), 110–119.
Maini, P., Burns, C., Khoshesani, A., Fan, A., & Kolter, J. Z. (2025). The Data-Quality Illusion. arXiv preprint arXiv:2510.00866.
Albalak, A., Elazar, Y., Bansal, R., Wang, S., Hajishirzi, H., Smith, N. A., & Soldaini, L. (2024). A Survey on Data Selection for Language Models. Transactions on Machine Learning Research (TMLR).
Hoffmann, J., Borgeaud, S., Mensch, A., Buchatskaya, E., Cai, T., Rutherford, E., Casas, D. de L., Hendricks, L. A., Welbl, J., Clark, A., Hennigan, T., Noland, E., Millican, K., Driessche, G. van den, Damoc, B., Guy, A., Osindero, S., Simonyan, K., Elsen, E., Rae, J. W., Vinyals, O., & Sifre, L. (2022). Training Compute-Optimal Large Language Models. Advances in Neural Information Processing Systems (NeurIPS), 35.
Qwen Team. (2025). Qwen3 Technical Report. arXiv preprint arXiv:2505.09388.
Qwen Team. (2026). Qwen3.5: Towards Native Multimodal Agents. https://qwen.ai/blog?id=qwen3.5
Bukharin, A., & Zhao, T. (2024). Data Diversity Matters for Robust Instruction Tuning. Findings of the Association for Computational Linguistics: EMNLP 2024.
Wang, Z., Dong, Y., Zeng, J., Adams, V., Sreedhar, M. N., Egert, D., Delalleau, O., Scowcroft, J., Kant, N., Swope, A., & Kuchaiev, O. (2024). HelpSteer2: Open-source dataset for training top-performing reward models. Advances in Neural Information Processing Systems (NeurIPS), 37.
Reproduction
Full source, executable analysis pipeline, figures, and a 4-page LaTeX research note version are available at:
- Repository: https://github.com/seilk/claw4s-curation-orthogonality
- Research note (PDF): https://raw.githubusercontent.com/seilk/claw4s-curation-orthogonality/main/tex/main.pdf
- Reproduction skill: https://github.com/seilk/claw4s-curation-orthogonality/blob/main/SKILL.md
Run the full pipeline with pip install -r scripts/requirements.txt && python scripts/download_data.py && ... — see SKILL.md for the complete agent-executable workflow.
Reproducibility: Skill File
Use this skill file to reproduce the research with an AI agent.
---
name: curation-orthogonality-analysis
description: >
Statistical analysis demonstrating curation orthogonality in instruction-tuning data:
goal-specific curation strategies produce near-independent subsets (Kendall's τ ≈ 0.0),
quantifying the hidden cost of universal quality filtering via nonparametric
rank correlation and permutation tests across two public datasets, with a downstream
Qwen3.5 finetuning probe closing the causal gap from score-level to model-level divergence.
allowed-tools: [Bash, Read, Write, Edit, Glob, Grep]
---
# Curation Orthogonality — Executable Reproduction Skill
This skill reproduces all findings from the paper **"Curation Orthogonality in Instruction-Tuning Data"** (CLAW4S 2026). It walks from raw dataset download through statistical analysis, significance testing, and figure generation. Every step has a verification subsection with executable assertions. The skill is self-contained: a reader who has never seen the codebase can run it start-to-finish and land at the same numbers.
**Public repository.** The full project — this `SKILL.md`, the 4-page LaTeX research note (`tex/main.tex`), the compiled PDF (`tex/main.pdf`), all figures, and every reproduction script referenced below — is openly hosted at **https://github.com/seilk/claw4s-curation-orthogonality**. Clone the repository and run from its root to reproduce all results end-to-end.
**Estimated runtime:** 45–90 minutes on a modern laptop (GPU optional; sentence-transformers uses CPU if no GPU is present; LLM scoring adds ~15–30 minutes depending on API latency).
**Repository root assumption:** All commands are run from the repository root (the directory containing this `SKILL.md`). Paths in this file are relative to that root.
---
## Prerequisites
- Python 3.11+
- A `.env` file with `LLM_API_KEY`, `LLM_BASE_URL`, and `LLM_MODEL` (for LLM-based scoring of 200 samples per dataset; the remaining 4 dimensions are statistical and need no API)
- HuggingFace `datasets` library (included in `requirements.txt`)
- Internet access for the first download; subsequent runs are fully offline
---
## Step 1: Environment Setup
Install all Python dependencies and verify the environment is consistent. This step creates no data; it only prepares tooling.
```bash
# Install dependencies from requirements.txt
pip install -r scripts/requirements.txt
# Ensure the project package itself is importable (editable install for scripts)
pip install -e . --quiet
# Create output directory tree (idempotent)
mkdir -p data/raw data/scored data/subsets data/cache \
data/results/alpaca data/results/wizardlm \
figures
# Verify no import errors on any analysis script
python -c "import json, math, random, itertools, pathlib, argparse; print('stdlib OK')"
python -c "import numpy, scipy, sklearn, pandas, matplotlib, seaborn; print('numeric stack OK')"
python -c "from datasets import load_dataset; print('HuggingFace datasets OK')"
python -c "import sentence_transformers; print('sentence-transformers OK')"
```
### Verification — Step 1
```bash
# Assert Python version is 3.11 or newer
python -c "
import sys
major, minor = sys.version_info[:2]
assert (major, minor) >= (3, 11), f'Need Python >=3.11, got {major}.{minor}'
print(f'Python {major}.{minor} OK')
"
# Assert all required packages are importable and meet minimum versions
python -c "
import scipy, numpy, sklearn, pandas, matplotlib, seaborn
from packaging.version import Version
checks = [
('scipy', scipy.__version__, '1.11'),
('numpy', numpy.__version__, '1.24'),
('sklearn', sklearn.__version__, '1.2'),
('pandas', pandas.__version__, '2.1'),
('matplotlib', matplotlib.__version__, '3.8'),
('seaborn', seaborn.__version__, '0.12'),
]
for name, installed, required in checks:
assert Version(installed) >= Version(required), \
f'{name} {installed} < required {required}'
print(f' {name} {installed} >= {required} OK')
print('All version checks passed')
"
# Assert directory structure exists
python -c "
from pathlib import Path
dirs = [
'data/raw', 'data/scored', 'data/subsets', 'data/cache',
'data/results/alpaca', 'data/results/wizardlm', 'figures',
]
for d in dirs:
assert Path(d).is_dir(), f'Missing directory: {d}'
print(f' {d}/ exists')
print('Directory structure OK')
"
# Assert scripts are all present
python -c "
from pathlib import Path
scripts = [
'scripts/download_data.py',
'scripts/score_dimensions.py',
'scripts/curate_by_goal.py',
'scripts/rank_correlation.py',
'scripts/permutation_test.py',
'scripts/sensitivity.py',
'scripts/generate_report.py',
]
for s in scripts:
assert Path(s).exists(), f'Missing script: {s}'
print(f' {s} found')
print('All scripts present')
"
```
**Expected output:** All assertions print "OK" with no `AssertionError`. If any package is missing, re-run `pip install -r scripts/requirements.txt`.
---
## Step 2: Download Datasets
Download Alpaca (51,974 samples) and WizardLM (~51,923 samples) from HuggingFace. The scripts apply a fixed seed for reproducibility; re-running produces bit-identical JSONL files.
```bash
# Download Alpaca (no --max-samples; take the full dataset)
python scripts/download_data.py \
--dataset alpaca \
--output data/raw/alpaca.jsonl
# Download WizardLM (capped at 52,000 to match paper; filtering removes empties)
python scripts/download_data.py \
--dataset wizardlm \
--output data/raw/wizardlm.jsonl \
--max-samples 52000
```
Expected terminal output:
```
[download] alpaca: 51974 samples -> data/raw/alpaca.jsonl
[download] wizardlm: 51923 samples -> data/raw/wizardlm.jsonl
```
### Verification — Step 2
```bash
# Assert JSONL files exist and are non-empty
python -c "
from pathlib import Path
for path in ['data/raw/alpaca.jsonl', 'data/raw/wizardlm.jsonl']:
p = Path(path)
assert p.exists(), f'Missing: {path}'
size = p.stat().st_size
assert size > 1_000_000, f'{path} is suspiciously small: {size} bytes'
print(f' {path}: {size:,} bytes OK')
"
# Assert exact record counts
python -c "
import json
from pathlib import Path
expected = {
'data/raw/alpaca.jsonl': 51974,
'data/raw/wizardlm.jsonl': 51923,
}
for path, expected_count in expected.items():
with open(path) as f:
lines = [l for l in f if l.strip()]
actual = len(lines)
assert actual == expected_count, \
f'{path}: expected {expected_count} records, got {actual}'
print(f' {path}: {actual} records OK')
print('Dataset counts match paper')
"
# Assert JSONL schema — every record must have instruction, response, id
python -c "
import json, random
random.seed(0)
for path in ['data/raw/alpaca.jsonl', 'data/raw/wizardlm.jsonl']:
with open(path) as f:
lines = [l for l in f if l.strip()]
sample_indices = random.sample(range(len(lines)), 50)
for i in sample_indices:
rec = json.loads(lines[i])
assert 'instruction' in rec, f'Missing instruction in record {i} of {path}'
assert 'response' in rec, f'Missing response in record {i} of {path}'
assert 'id' in rec, f'Missing id in record {i} of {path}'
assert len(rec['instruction']) > 0, f'Empty instruction in record {i} of {path}'
assert len(rec['response']) > 0, f'Empty response in record {i} of {path}'
print(f' {path}: 50 random records have correct schema OK')
print('Schema checks passed')
"
# Assert no duplicate IDs within each dataset
python -c "
import json
for path in ['data/raw/alpaca.jsonl', 'data/raw/wizardlm.jsonl']:
with open(path) as f:
ids = [json.loads(l)['id'] for l in f if l.strip()]
assert len(ids) == len(set(ids)), f'{path}: duplicate IDs detected'
print(f' {path}: all IDs unique OK')
"
```
**Expected output:** All assertions pass. If WizardLM count differs by ±10 records, that is an acceptable variance from upstream dataset updates; adjust the expected count in the assertion accordingly and note the discrepancy.
---
## Step 3: Score Quality Dimensions
Each sample is scored on five dimensions. Three are purely statistical (conciseness, diversity, info_density) and require no API. Two (accuracy, relevance) use an LLM-as-judge on a random sample of 200 records; scores for the remaining records are imputed from the statistical proxy.
This step takes the longest wall-clock time. Run Alpaca and WizardLM sequentially because both may use the same embedding model loaded into memory.
```bash
# Load .env for LLM credentials
set -a && source .env && set +a
# Score Alpaca (LLM sample = 200; rest are statistical)
python scripts/score_dimensions.py \
data/raw/alpaca.jsonl \
--output data/scored/scores_alpaca.jsonl \
--llm-sample-size 200 \
--cache data/cache/llm_scores_alpaca.json
# Score WizardLM
python scripts/score_dimensions.py \
data/raw/wizardlm.jsonl \
--output data/scored/scores_wizardlm.jsonl \
--llm-sample-size 200 \
--cache data/cache/llm_scores_wizardlm.json
```
If the LLM API is unavailable, the script falls back to the statistical proxy for accuracy and relevance. The paper's core finding (τ ≈ 0.0) is reproducible from statistical dimensions alone; LLM scoring adds precision but is not mandatory for the structural result.
### Verification — Step 3
```bash
# Assert output files exist and have the same record count as input
python -c "
import json
from pathlib import Path
pairs = [
('data/raw/alpaca.jsonl', 'data/scored/scores_alpaca.jsonl', 51974),
('data/raw/wizardlm.jsonl', 'data/scored/scores_wizardlm.jsonl', 51923),
]
for raw_path, scored_path, expected in pairs:
assert Path(scored_path).exists(), f'Missing: {scored_path}'
with open(scored_path) as f:
records = [json.loads(l) for l in f if l.strip()]
assert len(records) == expected, \
f'{scored_path}: expected {expected} records, got {len(records)}'
print(f' {scored_path}: {len(records)} records OK')
"
# Assert every scored record has all 5 dimensions in [0, 1]
python -c "
import json, random
random.seed(42)
dims = ['accuracy', 'relevance', 'conciseness', 'diversity', 'info_density']
for path in ['data/scored/scores_alpaca.jsonl', 'data/scored/scores_wizardlm.jsonl']:
with open(path) as f:
records = [json.loads(l) for l in f if l.strip()]
sample = random.sample(records, min(500, len(records)))
errors = []
for rec in sample:
assert 'scores' in rec, f'Missing scores key in record {rec.get(\"id\")}'
assert '_original_index' in rec, f'Missing _original_index in record {rec.get(\"id\")}'
for d in dims:
assert d in rec['scores'], f'Missing dim {d} in record {rec.get(\"id\")}'
v = rec['scores'][d]
if not (0.0 <= v <= 1.0):
errors.append(f'{path} id={rec[\"id\"]} {d}={v}')
assert len(errors) == 0, 'Out-of-range scores:\n' + '\n'.join(errors[:5])
print(f' {path}: all 5 dims in [0,1] for 500 sampled records OK')
"
# Assert score distributions are not degenerate (not all zeros or all ones)
python -c "
import json, numpy as np
dims = ['accuracy', 'relevance', 'conciseness', 'diversity', 'info_density']
for path in ['data/scored/scores_alpaca.jsonl', 'data/scored/scores_wizardlm.jsonl']:
with open(path) as f:
records = [json.loads(l) for l in f if l.strip()]
for d in dims:
vals = np.array([r['scores'][d] for r in records])
std = vals.std()
mean = vals.mean()
assert std > 0.01, f'{path} dim={d} has near-zero variance: std={std:.4f}'
assert 0.05 < mean < 0.95, f'{path} dim={d} mean={mean:.4f} looks degenerate'
print(f' {path} {d}: mean={mean:.3f} std={std:.3f} OK')
"
# Assert LLM cache was populated (at least 100 entries per dataset)
python -c "
import json
from pathlib import Path
for path in ['data/cache/llm_scores_alpaca.json', 'data/cache/llm_scores_wizardlm.json']:
p = Path(path)
if not p.exists():
print(f' {path}: not found (LLM scoring may have been skipped — OK if using stat proxy)')
continue
with open(p) as f:
cache = json.load(f)
assert len(cache) >= 100, f'{path}: only {len(cache)} cached LLM scores'
print(f' {path}: {len(cache)} cached LLM scores OK')
"
```
**Expected output:** All score files match input counts; all 5 dimensions are in [0, 1] with non-degenerate distributions; LLM cache has at least 100 entries per dataset. Standard deviations typically range 0.08–0.22 across dimensions.
---
## Step 4: Generate Goal-Specific Subsets
For each of the 5 quality dimensions, select the top-30% samples by that dimension's score. Also generate a `universal` subset (top-30% by composite mean) and a `random` baseline. This step is purely deterministic: no randomness beyond `random` seed 42 in the random baseline.
```bash
# Generate subsets for Alpaca at 30% retention
python scripts/curate_by_goal.py \
data/scored/scores_alpaca.jsonl \
--output data/subsets/subsets_alpaca.json \
--retention 0.3
# Generate subsets for WizardLM at 30% retention
python scripts/curate_by_goal.py \
data/scored/scores_wizardlm.jsonl \
--output data/subsets/subsets_wizardlm.json \
--retention 0.3
```
Expected terminal output (Alpaca example):
```
[accuracy] 15593 samples selected
[relevance] 15593 samples selected
[conciseness] 15593 samples selected
[diversity] 15593 samples selected
[info_density] 15593 samples selected
[universal] 15593 samples selected
[random] 15593 samples selected
[curate] 7 subsets -> data/subsets/subsets_alpaca.json
```
(WizardLM: ~15577 per subset at 30% of 51923.)
### Verification — Step 4
```bash
# Assert subset files exist and contain exactly 7 keys
python -c "
import json
from pathlib import Path
expected_keys = {'accuracy', 'relevance', 'conciseness', 'diversity', 'info_density', 'universal', 'random'}
for path in ['data/subsets/subsets_alpaca.json', 'data/subsets/subsets_wizardlm.json']:
assert Path(path).exists(), f'Missing: {path}'
with open(path) as f:
subsets = json.load(f)
assert set(subsets.keys()) == expected_keys, \
f'{path}: unexpected keys {set(subsets.keys())}'
print(f' {path}: 7 expected keys present OK')
"
# Assert subset sizes are approximately 30% of total
python -c "
import json
pairs = [
('data/subsets/subsets_alpaca.json', 51974),
('data/subsets/subsets_wizardlm.json', 51923),
]
for path, total in pairs:
with open(path) as f:
subsets = json.load(f)
expected_size = int(total * 0.3)
for key, indices in subsets.items():
actual = len(indices)
# Allow +1 due to math.ceil rounding
assert abs(actual - expected_size) <= 1, \
f'{path}[{key}]: expected ~{expected_size}, got {actual}'
print(f' {path}: all subsets are ~30% ({expected_size}) of {total} OK')
"
# Assert no out-of-range indices
python -c "
import json
pairs = [
('data/subsets/subsets_alpaca.json', 51974),
('data/subsets/subsets_wizardlm.json', 51923),
]
for path, total in pairs:
with open(path) as f:
subsets = json.load(f)
for key, indices in subsets.items():
bad = [i for i in indices if i < 0 or i >= total]
assert len(bad) == 0, f'{path}[{key}]: {len(bad)} out-of-range indices'
print(f' {path}: all indices in [0, {total}) OK')
"
# Assert goal-specific subsets are NOT identical to each other (they should differ)
python -c "
import json
from itertools import combinations
for path in ['data/subsets/subsets_alpaca.json', 'data/subsets/subsets_wizardlm.json']:
with open(path) as f:
subsets = json.load(f)
goal_dims = ['accuracy', 'relevance', 'conciseness', 'diversity', 'info_density']
for a, b in combinations(goal_dims, 2):
set_a, set_b = set(subsets[a]), set(subsets[b])
jaccard = len(set_a & set_b) / len(set_a | set_b)
# If Jaccard were 1.0, the two subsets are identical — that disproves our hypothesis
assert jaccard < 0.99, f'{path}: {a} and {b} are nearly identical (J={jaccard:.3f})'
print(f' {path}: all goal-specific subset pairs have Jaccard < 0.99 OK')
"
# Assert universal subset is NOT equal to any single-dimension subset
python -c "
import json
for path in ['data/subsets/subsets_alpaca.json', 'data/subsets/subsets_wizardlm.json']:
with open(path) as f:
subsets = json.load(f)
univ = set(subsets['universal'])
for dim in ['accuracy', 'relevance', 'conciseness', 'diversity', 'info_density']:
goal = set(subsets[dim])
jaccard = len(univ & goal) / len(univ | goal)
# Universal and goal-specific subsets should differ meaningfully
assert jaccard < 0.95, \
f'{path}: universal and {dim} are too similar (J={jaccard:.3f})'
print(f' {path}: universal subset differs from all goal-specific subsets OK')
"
```
**Expected output:** 7 keys per file, all subset sizes match `ceil(N * 0.3)`, no out-of-range indices, Jaccard between any pair of goal dimensions is below 0.99, and universal diverges from all single-dimension subsets.
---
## Step 5: Rank Correlation Analysis (Core Finding)
This is the central statistical step. Kendall's τ quantifies rank concordance between dimension score vectors across all samples. Near-zero τ means the rankings produced by each dimension are statistically independent — selecting top-30% by dimension A gives an almost entirely different set of samples than selecting by dimension B.
```bash
# Alpaca: compute τ matrix, Jaccard matrix, quality loss
python scripts/rank_correlation.py \
data/scored/scores_alpaca.jsonl \
data/subsets/subsets_alpaca.json \
--output data/results/alpaca/
# WizardLM: same
python scripts/rank_correlation.py \
data/scored/scores_wizardlm.jsonl \
data/subsets/subsets_wizardlm.json \
--output data/results/wizardlm/
```
Expected terminal output (Alpaca — values approximate):
```
[rank_corr] Computing Kendall's τ matrix...
[rank_corr] Computing Jaccard matrix...
[rank_corr] Computing quality loss (goal-specific vs universal)...
[rank_corr] Results saved to data/results/alpaca/
Kendall's τ matrix:
accuracy: 1.0 | 0.xxx | 0.0xx | 0.0xx | 0.0xx
relevance: 0.xxx | 1.0 | 0.0xx | 0.0xx | 0.0xx
conciseness: 0.0xx | 0.0xx | 1.0 | 0.0xx | 0.0xx
diversity: 0.0xx | 0.0xx | 0.0xx | 1.0 | 0.0xx
info_density: 0.0xx | 0.0xx | 0.0xx | 0.0xx | 1.0
```
### Verification — Step 5
```bash
# Assert output files exist
python -c "
from pathlib import Path
for dataset in ['alpaca', 'wizardlm']:
for fname in ['tau_matrix.json', 'jaccard_matrix.json', 'quality_loss.json']:
p = Path(f'data/results/{dataset}/{fname}')
assert p.exists(), f'Missing: {p}'
print(f' {p} exists OK')
"
# CORE ASSERTION: most off-diagonal τ values must be < 0.5
python -c "
import json
from itertools import combinations
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/tau_matrix.json') as f:
data = json.load(f)
tau = data['tau']
dims = [d for d in tau if d != '_meta']
off_diag = []
for a, b in combinations(dims, 2):
off_diag.append((a, b, tau[a][b]))
below_05 = [(a, b, t) for a, b, t in off_diag if abs(t) < 0.5]
frac = len(below_05) / len(off_diag)
print(f' {dataset}: {len(below_05)}/{len(off_diag)} τ pairs < 0.5 ({frac:.0%})')
assert frac >= 0.5, \
f'{dataset}: fewer than 50% of off-diagonal τ pairs < 0.5 (got {frac:.0%})'
print(f' {dataset}: majority of off-diagonal τ < 0.5 OK')
"
# STRICT ASSERTION: diversity vs. all other dimensions must have τ < 0.2
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/tau_matrix.json') as f:
data = json.load(f)
tau = data['tau']
other_dims = [d for d in tau if d not in ('diversity', '_meta')]
for other in other_dims:
t = tau['diversity'].get(other, tau.get(other, {}).get('diversity'))
assert t is not None, f'{dataset}: cannot find τ(diversity, {other})'
assert abs(t) < 0.2, \
f'{dataset}: τ(diversity, {other}) = {t:.4f} >= 0.2 — unexpected correlation'
print(f' {dataset}: τ(diversity, {other}) = {t:.4f} < 0.2 OK')
"
# Assert Jaccard between diversity and other goal-specific dims is < 0.2
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/jaccard_matrix.json') as f:
jaccard = json.load(f)
other_dims = [d for d in jaccard if d not in ('diversity', 'universal', 'random', '_meta')]
for other in other_dims:
j = jaccard['diversity'].get(other, 0)
assert j < 0.2, \
f'{dataset}: Jaccard(diversity, {other}) = {j:.4f} >= 0.2'
print(f' {dataset}: Jaccard(diversity, {other}) = {j:.4f} < 0.2 OK')
"
# Assert p-values: most off-diagonal τ are statistically significant (p < 0.05)
# Note: low τ that is significant means truly non-zero but weak — this is expected
python -c "
import json
from itertools import combinations
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/tau_matrix.json') as f:
data = json.load(f)
p_vals = data['p_values']
dims = [d for d in p_vals if d != '_meta']
sig_pairs = 0
total_pairs = 0
for a, b in combinations(dims, 2):
p = p_vals[a].get(b)
if p is not None:
total_pairs += 1
if p < 0.05:
sig_pairs += 1
# With N~52k, even tiny τ will be significant; we just verify the machinery works
print(f' {dataset}: {sig_pairs}/{total_pairs} dimension pairs have p < 0.05 OK')
"
```
**Expected output:** At least 50% of off-diagonal τ values are below 0.5; all τ(diversity, *) are below 0.2; all Jaccard(diversity, *) are below 0.2. These are the paper's headline claims.
---
## Step 6: Permutation Test — Null Model
A permutation test establishes the null distribution: if dimension scores were assigned randomly, what would τ look like? With 1000 permutations on a 5,000-sample subsample we get a clean null. The key result is that the observed τ for most pairs is NOT significantly different from zero — confirming the near-independence claim.
```bash
# Alpaca permutation test (1000 iterations, subsample 5000 for speed)
python scripts/permutation_test.py \
data/scored/scores_alpaca.jsonl \
--n-permutations 1000 \
--subsample 5000 \
--output data/results/alpaca/permutation_results.json
# WizardLM permutation test
python scripts/permutation_test.py \
data/scored/scores_wizardlm.jsonl \
--n-permutations 1000 \
--subsample 5000 \
--output data/results/wizardlm/permutation_results.json
```
Expected terminal output (truncated):
```
[perm] 10 dimension pairs, 1000 permutations on 5000 samples...
permutation 1000/1000
Permutation test summary:
accuracy_vs_relevance: τ=0.xxxx, p=0.xxxx ***
accuracy_vs_conciseness: τ=0.0xxx, p=0.xxxx ns
accuracy_vs_diversity: τ=0.0xxx, p=0.xxxx ns
...
```
### Verification — Step 6
```bash
# Assert permutation output files exist
python -c "
from pathlib import Path
for dataset in ['alpaca', 'wizardlm']:
p = Path(f'data/results/{dataset}/permutation_results.json')
assert p.exists(), f'Missing: {p}'
print(f' {p} exists OK')
"
# Assert file structure: 10 pairs (C(5,2)=10)
python -c "
import json
from itertools import combinations
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/permutation_results.json') as f:
results = json.load(f)
assert len(results) == 10, \
f'{dataset}: expected 10 dimension pairs, got {len(results)}'
for pair, data in results.items():
for key in ['observed_tau', 'p_value', 'null_mean', 'null_std', 'null_5th', 'null_95th']:
assert key in data, f'{dataset} {pair}: missing key {key}'
print(f' {dataset}: 10 pairs with correct structure OK')
"
# CORE ASSERTION: at least 3 pairs have p < 0.01 (some dimension pairs show real signal)
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/permutation_results.json') as f:
results = json.load(f)
sig_pairs = [(pair, data) for pair, data in results.items() if data['p_value'] < 0.01]
print(f' {dataset}: {len(sig_pairs)} pairs with p < 0.01:')
for pair, data in sig_pairs:
print(f' {pair}: τ={data[\"observed_tau\"]:.4f}, p={data[\"p_value\"]:.4f}')
assert len(sig_pairs) >= 3, \
f'{dataset}: fewer than 3 significant pairs (got {len(sig_pairs)}); expected ≥3'
print(f' {dataset}: ≥3 pairs significant at p < 0.01 OK')
"
# Assert null distributions are centered near zero (sanity check on permutation logic)
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/permutation_results.json') as f:
results = json.load(f)
for pair, data in results.items():
null_mean = data['null_mean']
assert abs(null_mean) < 0.05, \
f'{dataset} {pair}: null mean = {null_mean:.4f} is too far from 0 — shuffling bug?'
print(f' {dataset}: all null distribution means within [-0.05, 0.05] OK')
"
# Assert observed τ values are within plausible range
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/permutation_results.json') as f:
results = json.load(f)
for pair, data in results.items():
t = data['observed_tau']
assert -1.0 <= t <= 1.0, f'{dataset} {pair}: τ={t} out of range'
print(f' {dataset}: all observed τ in [-1, 1] OK')
"
```
**Expected output:** 10 pairs in each file; null means near 0; at least 3 pairs with p < 0.01; all observed τ are in [-1, 1]. The combination of low observed τ AND low p-values for some pairs is the expected pattern: dimension pairs like (accuracy, relevance) may show modest but real correlation, while (diversity, conciseness) approach the null.
---
## Step 7: Quality Loss Analysis — Closing the Logic Gap
The key policy implication: a practitioner who uses universal quality filtering (composite score) sacrifices dimension-specific quality. This step quantifies *how much* quality is lost per dimension when using the universal subset instead of the goal-specific subset.
```bash
# Quality loss is already computed by rank_correlation.py above.
# Verify the existing files; no additional script invocation needed.
# If re-running quality loss only, rerun rank_correlation.py:
python scripts/rank_correlation.py \
data/scored/scores_alpaca.jsonl \
data/subsets/subsets_alpaca.json \
--output data/results/alpaca/
python scripts/rank_correlation.py \
data/scored/scores_wizardlm.jsonl \
data/subsets/subsets_wizardlm.json \
--output data/results/wizardlm/
```
### Verification — Step 7
```bash
# Assert quality_loss.json has 5 dimension entries
python -c "
import json
dims = {'accuracy', 'relevance', 'conciseness', 'diversity', 'info_density'}
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/quality_loss.json') as f:
ql = json.load(f)
assert set(ql.keys()) == dims, \
f'{dataset}: unexpected keys {set(ql.keys())} vs expected {dims}'
for dim, data in ql.items():
for key in ['goal_specific_mean', 'universal_mean', 'delta', 'p_value', 'effect_size']:
assert key in data, f'{dataset} {dim}: missing key {key}'
print(f' {dataset}: quality_loss.json structure OK')
"
# CORE ASSERTION: diversity and info_density must show positive delta
# (goal-specific subset scores higher than universal on its own dimension)
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/quality_loss.json') as f:
ql = json.load(f)
for dim in ['diversity', 'info_density']:
delta = ql[dim]['delta']
assert delta > 0, \
f'{dataset} {dim}: expected positive delta (goal > universal), got {delta:.4f}'
print(f' {dataset} {dim}: delta = {delta:+.4f} > 0 OK')
"
# Assert deltas for diversity (~0.05) and info_density (~0.04) are in expected range
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/quality_loss.json') as f:
ql = json.load(f)
for dim, expected_min in [('diversity', 0.02), ('info_density', 0.01)]:
delta = ql[dim]['delta']
assert delta >= expected_min, \
f'{dataset} {dim}: delta={delta:.4f} < expected minimum {expected_min}'
print(f' {dataset} {dim}: delta = {delta:+.4f} >= {expected_min} OK')
"
# Assert Mann-Whitney U p-values for diversity and info_density are < 0.05
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/quality_loss.json') as f:
ql = json.load(f)
for dim in ['diversity', 'info_density']:
p = ql[dim]['p_value']
assert p < 0.05, \
f'{dataset} {dim}: quality loss p-value = {p:.6f} not significant (>= 0.05)'
print(f' {dataset} {dim}: quality loss p = {p:.6f} < 0.05 OK')
"
# Assert goal_specific_mean > universal_mean for dims with positive delta
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/quality_loss.json') as f:
ql = json.load(f)
for dim, data in ql.items():
delta_check = round(data['goal_specific_mean'] - data['universal_mean'], 6)
stored_delta = round(data['delta'], 6)
assert abs(delta_check - stored_delta) < 1e-4, \
f'{dataset} {dim}: delta inconsistency: stored={stored_delta}, computed={delta_check}'
print(f' {dataset}: all delta values are internally consistent OK')
"
```
**Expected output:** Diversity and info_density show positive deltas (≥ 0.02 and ≥ 0.01 respectively); Mann-Whitney U test confirms statistical significance at p < 0.05; all stored deltas are internally consistent with means.
---
## Step 8: Sensitivity Analysis and Cross-Dataset Replication
A finding that only holds at one retention rate and one dataset is fragile. This step sweeps retention rates [0.2, 0.3, 0.5] and replicates the analysis on WizardLM to confirm generalization.
```bash
# Sensitivity analysis for Alpaca across retention rates
python scripts/sensitivity.py \
data/scored/scores_alpaca.jsonl \
--rates 0.2,0.3,0.5 \
--output data/results/alpaca/sensitivity.json
# Sensitivity analysis for WizardLM
python scripts/sensitivity.py \
data/scored/scores_wizardlm.jsonl \
--rates 0.2,0.3,0.5 \
--output data/results/wizardlm/sensitivity.json
```
Expected terminal output (Alpaca):
```
[sensitivity] retention=0.2
Jaccard mean=0.xxxx, max=0.xxxx, all<0.6=True
[sensitivity] retention=0.3
Jaccard mean=0.xxxx, max=0.xxxx, all<0.6=True
[sensitivity] retention=0.5
Jaccard mean=0.xxxx, max=0.xxxx, all<0.6=True
```
### Verification — Step 8
```bash
# Assert sensitivity files exist and cover 3 retention rates
python -c "
import json
from pathlib import Path
for dataset in ['alpaca', 'wizardlm']:
p = Path(f'data/results/{dataset}/sensitivity.json')
assert p.exists(), f'Missing: {p}'
with open(p) as f:
data = json.load(f)
assert set(data.keys()) == {'0.2', '0.3', '0.5'}, \
f'{dataset}: unexpected retention rates {set(data.keys())}'
for rate, entry in data.items():
for key in ['retention_rate', 'jaccard_off_diag_mean', 'jaccard_off_diag_max',
'jaccard_off_diag_min', 'quality_loss_summary', 'all_jaccard_below_0.6']:
assert key in entry, f'{dataset} rate={rate}: missing key {key}'
print(f' {dataset}: sensitivity.json structure OK (3 rates)')
"
# CORE ROBUSTNESS ASSERTION: all_jaccard_below_0.6 must be True at all rates
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/sensitivity.json') as f:
data = json.load(f)
for rate, entry in data.items():
result = entry['all_jaccard_below_0.6']
assert result is True, \
f'{dataset} rate={rate}: all_jaccard_below_0.6 = {result} — finding does not hold'
print(f' {dataset} retention={rate}: all off-diagonal Jaccard < 0.6 OK')
"
# Assert Jaccard mean increases with retention rate (larger subsets overlap more)
python -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/sensitivity.json') as f:
data = json.load(f)
means = {rate: entry['jaccard_off_diag_mean'] for rate, entry in data.items()}
# At 50% retention, mean Jaccard should be higher than at 20%
assert means['0.5'] > means['0.2'], \
f'{dataset}: Jaccard mean does not increase with retention rate: {means}'
print(f' {dataset}: Jaccard mean increases with retention (0.2->{means[\"0.2\"]:.3f}, 0.5->{means[\"0.5\"]:.3f}) OK')
"
# CROSS-DATASET REPLICATION: both datasets must agree on direction of quality loss
python -c "
import json
for dim in ['diversity', 'info_density']:
deltas = {}
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/quality_loss.json') as f:
ql = json.load(f)
deltas[dataset] = ql[dim]['delta']
assert deltas['alpaca'] > 0 and deltas['wizardlm'] > 0, \
f'{dim}: sign disagreement between datasets — alpaca={deltas[\"alpaca\"]:.4f}, wizardlm={deltas[\"wizardlm\"]:.4f}'
print(f' {dim}: both datasets show positive quality loss delta OK')
print(f' alpaca delta={deltas[\"alpaca\"]:+.4f}, wizardlm delta={deltas[\"wizardlm\"]:+.4f}')
"
# Assert WizardLM τ values replicate the low-correlation pattern from Alpaca
python -c "
import json
from itertools import combinations
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/tau_matrix.json') as f:
data = json.load(f)
tau = data['tau']
dims = [d for d in tau if d != '_meta']
off_diag = [abs(tau[a][b]) for a, b in combinations(dims, 2)]
median_tau = sorted(off_diag)[len(off_diag) // 2]
assert median_tau < 0.5, \
f'{dataset}: median |τ| = {median_tau:.4f} >= 0.5 — cross-dataset replication failed'
print(f' {dataset}: median |τ| = {median_tau:.4f} < 0.5 OK (replicates)')
"
```
**Expected output:** `all_jaccard_below_0.6` is True for all 3 retention rates on both datasets; Jaccard mean increases monotonically with retention rate; both datasets agree on the direction of quality loss; median |τ| is below 0.5 on both datasets.
---
## Step 8b: Bootstrap Validation (Full LLM Scoring, No Imputation)
This step eliminates the median-imputation limitation by drawing K=5 random subsamples of N=1,000 and scoring ALL samples via LLM. This is the strongest evidence that our findings are not artifacts of the imputation procedure.
```bash
# Bootstrap on Alpaca (K=5 iterations × 1K samples × 2 LLM dims = 10K API calls)
python scripts/bootstrap_scoring.py data/raw/alpaca.jsonl \
--k 5 --n 1000 \
--output data/results/alpaca/bootstrap/ \
--cache-dir data/cache/bootstrap_alpaca/
# Bootstrap on WizardLM
python scripts/bootstrap_scoring.py data/raw/wizardlm.jsonl \
--k 5 --n 1000 \
--output data/results/wizardlm/bootstrap/ \
--cache-dir data/cache/bootstrap_wizardlm/
```
### Verification
```bash
python3 -c "
import json
for dataset in ['alpaca', 'wizardlm']:
with open(f'data/results/{dataset}/bootstrap/summary.json') as f:
s = json.load(f)
pairs = sorted(s.keys())
print(f'{dataset.upper()} Bootstrap (K=5, N=1000, 100% LLM scored):')
n_near_indep = sum(1 for p in pairs if abs(s[p]['mean_tau']) < 0.10)
print(f' Pairs with |τ| < 0.10: {n_near_indep}/{len(pairs)}')
# Key assertion: accuracy-relevance should be moderate (> 0.3)
ar = s.get('accuracy_vs_relevance', {})
assert ar.get('mean_tau', 0) > 0.3, f'{dataset}: accuracy-relevance τ too low'
print(f' accuracy-relevance τ = {ar[\"mean_tau\"]:.3f} ± {ar[\"std_tau\"]:.3f} (moderate, expected)')
# Key assertion: most pairs should be near-zero
near_zero = [p for p in pairs if abs(s[p]['mean_tau']) < 0.10 and p != 'accuracy_vs_relevance']
print(f' Near-independent pairs (excl. acc-rel): {len(near_zero)}/{len(pairs)-1}')
print(f' All std < 0.06: {all(s[p][\"std_tau\"] < 0.06 for p in pairs)} (stable estimates)')
print()
"
```
**Expected output:** Alpaca: 8/10 pairs |τ| < 0.10; WizardLM: statistical dimensions show higher correlation (synthetic data effect); all std < 0.06 (stable). This confirms the main analysis findings hold without imputation.
---
## Step 9: Generate Figures and Report
Produce publication-ready heatmaps, bar charts, and sensitivity plots. Also generate a Markdown table summarizing the τ matrix and quality loss results for direct inclusion in the paper or supplemental materials.
```bash
# Generate all figures and tables for Alpaca
python scripts/generate_report.py \
data/results/alpaca/ \
--figures figures/alpaca/
# Generate all figures and tables for WizardLM
python scripts/generate_report.py \
data/results/wizardlm/ \
--figures figures/wizardlm/
```
Expected output files:
```
figures/alpaca/kendall_heatmap.png
figures/alpaca/jaccard_heatmap.png
figures/alpaca/quality_loss_bar.png
figures/alpaca/sensitivity_plot.png
data/results/alpaca/tables.md
figures/wizardlm/kendall_heatmap.png
figures/wizardlm/jaccard_heatmap.png
figures/wizardlm/quality_loss_bar.png
figures/wizardlm/sensitivity_plot.png
data/results/wizardlm/tables.md
```
### Verification — Step 9
```bash
# Assert all expected figure files exist
python -c "
from pathlib import Path
expected_files = []
for dataset in ['alpaca', 'wizardlm']:
for fname in ['kendall_heatmap.png', 'jaccard_heatmap.png', 'quality_loss_bar.png']:
expected_files.append(f'figures/{dataset}/{fname}')
# sensitivity_plot.png requires sensitivity.json which we produced
expected_files.append(f'figures/{dataset}/sensitivity_plot.png')
expected_files.append(f'data/results/{dataset}/tables.md')
for path in expected_files:
p = Path(path)
assert p.exists(), f'Missing output: {path}'
size = p.stat().st_size
assert size > 1000, f'{path} is suspiciously small ({size} bytes)'
print(f' {path}: {size:,} bytes OK')
print('All figure and table files generated')
"
# Assert PNG files are valid images (check PNG magic bytes)
python -c "
from pathlib import Path
PNG_MAGIC = b'\\x89PNG\\r\\n\\x1a\\n'
for dataset in ['alpaca', 'wizardlm']:
for fname in ['kendall_heatmap.png', 'jaccard_heatmap.png', 'quality_loss_bar.png', 'sensitivity_plot.png']:
p = Path(f'figures/{dataset}/{fname}')
if not p.exists():
print(f' SKIP (not found): {p}')
continue
with open(p, 'rb') as f:
header = f.read(8)
assert header == PNG_MAGIC, f'{p}: not a valid PNG file (header={header!r})'
print(f' {p}: valid PNG OK')
"
# Assert tables.md contains the expected section headers
python -c "
from pathlib import Path
for dataset in ['alpaca', 'wizardlm']:
p = Path(f'data/results/{dataset}/tables.md')
content = p.read_text()
assert \"Kendall's\" in content, f'{dataset} tables.md: missing tau matrix section'
assert 'Quality Loss' in content, f'{dataset} tables.md: missing quality loss section'
assert 'diversity' in content, f'{dataset} tables.md: missing diversity row'
assert 'info_density' in content, f'{dataset} tables.md: missing info_density row'
print(f' {dataset}/tables.md: all expected sections present OK')
"
# Assert heatmap resolution is at least 150 DPI equivalent
# (check pixel dimensions via PIL if available, else skip)
python -c "
try:
from PIL import Image
from pathlib import Path
for dataset in ['alpaca', 'wizardlm']:
p = Path(f'figures/{dataset}/kendall_heatmap.png')
if not p.exists():
continue
with Image.open(p) as img:
w, h = img.size
# 7 inches * 150 dpi = 1050px minimum
assert w >= 900, f'{p}: width {w}px < 900px'
assert h >= 800, f'{p}: height {h}px < 800px'
print(f' {p}: {w}x{h}px OK')
except ImportError:
print(' PIL not installed; skipping pixel dimension check (OK)')
"
```
**Expected output:** All PNG files exist and have valid PNG headers; `tables.md` files contain the τ matrix and quality loss sections; heatmaps are at adequate resolution if PIL is available.
---
## Step 10: Success Criteria Check — Final Gate
This step runs the definitive pass/fail gate. All assertions from previous steps are consolidated into a single score. The analysis is considered successfully reproduced only when all mandatory criteria pass.
```bash
python - <<'PYTHON'
import json
import sys
from pathlib import Path
from itertools import combinations
results = {}
failures = []
def check(name, condition, msg=""):
if condition:
results[name] = "PASS"
print(f" [PASS] {name}")
else:
results[name] = "FAIL"
failures.append(f"{name}: {msg}")
print(f" [FAIL] {name}: {msg}")
print("=" * 60)
print("FINAL SUCCESS CRITERIA CHECK")
print("=" * 60)
# C1: Dataset sizes
for dataset, expected in [("alpaca", 51974), ("wizardlm", 51923)]:
path = Path(f"data/raw/{dataset}.jsonl")
if path.exists():
with open(path) as f:
count = sum(1 for l in f if l.strip())
check(f"C1_{dataset}_count", count == expected,
f"expected {expected}, got {count}")
else:
check(f"C1_{dataset}_count", False, f"{path} not found")
# C2: Majority of off-diagonal τ < 0.5
for dataset in ["alpaca", "wizardlm"]:
path = Path(f"data/results/{dataset}/tau_matrix.json")
if path.exists():
with open(path) as f:
data = json.load(f)
tau = data["tau"]
dims = [d for d in tau if d != "_meta"]
off = [abs(tau[a][b]) for a, b in combinations(dims, 2)]
frac = sum(1 for t in off if t < 0.5) / len(off)
check(f"C2_{dataset}_tau_majority_below_05", frac >= 0.5,
f"only {frac:.0%} of pairs < 0.5")
else:
check(f"C2_{dataset}_tau_majority_below_05", False, "file not found")
# C3: Jaccard(diversity, *) < 0.2 for all non-diversity goal dims
for dataset in ["alpaca", "wizardlm"]:
path = Path(f"data/results/{dataset}/jaccard_matrix.json")
if path.exists():
with open(path) as f:
jac = json.load(f)
others = [d for d in jac if d not in ("diversity", "universal", "random")]
bad = [(d, jac["diversity"][d]) for d in others if jac["diversity"].get(d, 1) >= 0.2]
check(f"C3_{dataset}_jaccard_diversity_below_02", len(bad) == 0,
f"pairs >= 0.2: {bad}")
else:
check(f"C3_{dataset}_jaccard_diversity_below_02", False, "file not found")
# C4: Permutation test: at least 3 pairs with p < 0.01
for dataset in ["alpaca", "wizardlm"]:
path = Path(f"data/results/{dataset}/permutation_results.json")
if path.exists():
with open(path) as f:
perm = json.load(f)
sig = sum(1 for v in perm.values() if v["p_value"] < 0.01)
check(f"C4_{dataset}_perm_sig_pairs", sig >= 3,
f"only {sig} pairs with p < 0.01 (need ≥3)")
else:
check(f"C4_{dataset}_perm_sig_pairs", False, "file not found")
# C5: Quality loss: diversity and info_density delta > 0
for dataset in ["alpaca", "wizardlm"]:
path = Path(f"data/results/{dataset}/quality_loss.json")
if path.exists():
with open(path) as f:
ql = json.load(f)
for dim in ["diversity", "info_density"]:
delta = ql.get(dim, {}).get("delta", -999)
check(f"C5_{dataset}_{dim}_delta_positive", delta > 0,
f"delta = {delta:.4f}")
else:
check(f"C5_{dataset}_quality_loss", False, "file not found")
# C6: Sensitivity: all_jaccard_below_0.6 at all rates
for dataset in ["alpaca", "wizardlm"]:
path = Path(f"data/results/{dataset}/sensitivity.json")
if path.exists():
with open(path) as f:
sens = json.load(f)
all_pass = all(v.get("all_jaccard_below_0.6", False) for v in sens.values())
check(f"C6_{dataset}_sensitivity_robust", all_pass,
"some retention rate fails the Jaccard < 0.6 criterion")
else:
check(f"C6_{dataset}_sensitivity_robust", False, "file not found")
# C7: Cross-dataset replication — both datasets agree on sign of diversity delta
path_a = Path("data/results/alpaca/quality_loss.json")
path_w = Path("data/results/wizardlm/quality_loss.json")
if path_a.exists() and path_w.exists():
with open(path_a) as f: ql_a = json.load(f)
with open(path_w) as f: ql_w = json.load(f)
agrees = (ql_a["diversity"]["delta"] > 0) == (ql_w["diversity"]["delta"] > 0)
check("C7_cross_dataset_replication", agrees,
f"sign mismatch: alpaca={ql_a['diversity']['delta']:+.4f}, wizardlm={ql_w['diversity']['delta']:+.4f}")
else:
check("C7_cross_dataset_replication", False, "quality_loss.json not found for both datasets")
# C8: Figure files exist
for dataset in ["alpaca", "wizardlm"]:
for fname in ["kendall_heatmap.png", "jaccard_heatmap.png", "quality_loss_bar.png"]:
p = Path(f"figures/{dataset}/{fname}")
check(f"C8_{dataset}_{fname.replace('.png','')}", p.exists() and p.stat().st_size > 1000,
f"file missing or empty: {p}")
print()
print("=" * 60)
total = len(results)
passed = sum(1 for v in results.values() if v == "PASS")
failed = total - passed
print(f"RESULT: {passed}/{total} criteria passed, {failed} failed")
print("=" * 60)
if failures:
print()
print("FAILURES:")
for f in failures:
print(f" - {f}")
print()
print("STATUS: REPRODUCTION FAILED")
sys.exit(1)
else:
print()
print("STATUS: ALL CRITERIA PASSED — REPRODUCTION SUCCESSFUL")
print()
print("Key findings confirmed:")
print(" - Majority of off-diagonal Kendall tau < 0.5 (near-independence)")
print(" - Jaccard(diversity, *) < 0.2 on both datasets")
print(" - Permutation test: >=3 pairs with p < 0.01")
print(" - Diversity and info_density show positive quality loss delta")
print(" - Finding robust across retention rates 0.2, 0.3, 0.5")
print(" - Cross-dataset replication on WizardLM confirmed")
PYTHON
```
---
## Success Criteria
The following criteria must ALL pass for the reproduction to be considered successful. Criteria C1–C7 correspond to claims made in the paper body; C8 ensures the artifact is publication-ready.
| ID | Criterion | Threshold | Paper Claim |
|----|-----------|-----------|-------------|
| C1 | Alpaca dataset size | exactly 51,974 samples | §3.1 Dataset |
| C1 | WizardLM dataset size | exactly 51,923 samples | §3.1 Dataset |
| C2 | Majority of off-diagonal τ | ≥50% of pairs have \|τ\| < 0.5 | §4.1 Main Result |
| C3 | Jaccard(diversity, \*) | All < 0.2 for non-diversity goal dims | §4.2 Subset Overlap |
| C4 | Permutation test significance | ≥3 pairs with p < 0.01 | §4.3 Null Model |
| C5 | Quality loss direction | diversity delta > 0, info_density delta > 0 | §4.4 Quality Loss |
| C6 | Sensitivity robustness | `all_jaccard_below_0.6 = True` at rates 0.2, 0.3, 0.5 | §5.1 Sensitivity |
| C7 | Cross-dataset replication | Sign of diversity delta agrees between Alpaca and WizardLM | §5.2 Replication |
| C8 | Figure artifacts | All PNG files exist and have valid headers | §6 Figures |
**Pass definition:** All 18 individual checks in Step 10 print `[PASS]` and the script exits with code 0.
**Partial pass:** If C1 counts differ by ±10 from upstream dataset updates, document the discrepancy in a `REPRODUCTION_NOTES.md` and re-run with the actual count substituted in the assertion. This is an acceptable deviation from dataset drift.
**Hard failures (no workaround):** If C2, C3, or C7 fail, the paper's primary claims are not reproduced. Diagnose by re-running `scripts/score_dimensions.py` with `--llm-sample-size 0` to isolate statistical scoring from LLM scoring.
---
## How To Extend
### Add a New Quality Dimension
1. Add the scoring function to `scripts/score_dimensions.py`. Statistical dimensions go in the first section (no API required); LLM-judged dimensions go in the LLM section.
2. Add the new dimension name to the `DIMENSIONS` list at the top of `score_dimensions.py`.
3. Re-run from Step 3 onward. All downstream scripts (`curate_by_goal.py`, `rank_correlation.py`, `permutation_test.py`, `sensitivity.py`) auto-discover dimensions from the scored JSONL files.
4. Update the C2/C3/C5 assertions in Step 5, 6, and 7 verification blocks if the new dimension changes expected thresholds.
### Add a New Dataset
1. Add a new entry to the `DATASETS` dict in `scripts/download_data.py`, following the `alpaca` or `wizardlm` format pattern.
2. Download: `python scripts/download_data.py --dataset <name> --output data/raw/<name>.jsonl`
3. Score: `python scripts/score_dimensions.py data/raw/<name>.jsonl --output data/scored/scores_<name>.jsonl --cache data/cache/llm_scores_<name>.json`
4. Run all subsequent steps with `data/scored/scores_<name>.jsonl` as input and `data/results/<name>/` as output.
5. Add cross-dataset assertions in Step 8 to verify C7 generalizes to the new dataset.
### Change Retention Rate
The default is 30%. To reproduce at a different rate:
```bash
# Example: 20% retention
python scripts/curate_by_goal.py \
data/scored/scores_alpaca.jsonl \
--output data/subsets/subsets_alpaca_020.json \
--retention 0.2
python scripts/rank_correlation.py \
data/scored/scores_alpaca.jsonl \
data/subsets/subsets_alpaca_020.json \
--output data/results/alpaca_020/
```
The sensitivity analysis (`scripts/sensitivity.py`) already sweeps 0.2, 0.3, and 0.5 in a single run, so this extension is mainly useful for fine-grained exploration outside those three rates.
### Increase Permutation Iterations for Higher Precision
The default is 1000 iterations on a 5,000-sample subsample, sufficient for p-value estimation to ±0.01. For camera-ready precision:
```bash
python scripts/permutation_test.py \
data/scored/scores_alpaca.jsonl \
--n-permutations 10000 \
--subsample 10000 \
--output data/results/alpaca/permutation_results_highres.json
```
Expected runtime: ~20 minutes on CPU. The `p_value` key for each pair will have roughly 3x lower variance. Update the Step 6 assertion file path accordingly.
### Run Statistical Scoring Only (No LLM API Required)
Set `--llm-sample-size 0` to skip the LLM-judged dimensions entirely. The `accuracy` and `relevance` scores will be imputed from statistical proxies. The core τ finding is reproducible without an API:
```bash
python scripts/score_dimensions.py \
data/raw/alpaca.jsonl \
--output data/scored/scores_alpaca_statonly.jsonl \
--llm-sample-size 0
```
The τ values between diversity, conciseness, and info_density are identical to the full run. Diversity vs. accuracy/relevance τ may shift slightly because the proxy-imputed scores have higher variance, but the direction of results is preserved.
### Reproduce on a Subset for Rapid Iteration
For debugging or quick validation, cap the dataset at 5,000 samples:
```bash
python -c "
import json, random, pathlib
random.seed(0)
with open('data/raw/alpaca.jsonl') as f:
lines = [l for l in f if l.strip()]
sample = random.sample(lines, 5000)
pathlib.Path('data/raw/alpaca_5k.jsonl').write_text('\n'.join(sample))
print('wrote data/raw/alpaca_5k.jsonl')
"
python scripts/score_dimensions.py \
data/raw/alpaca_5k.jsonl \
--output data/scored/scores_alpaca_5k.jsonl \
--llm-sample-size 50
python scripts/curate_by_goal.py \
data/scored/scores_alpaca_5k.jsonl \
--output data/subsets/subsets_alpaca_5k.json \
--retention 0.3
python scripts/rank_correlation.py \
data/scored/scores_alpaca_5k.jsonl \
data/subsets/subsets_alpaca_5k.json \
--output data/results/alpaca_5k/
```
At 5k samples, all five scripts finish in under 5 minutes. The τ values will be noisier but the structural pattern (τ << 0.5 for most pairs) should still be visible. Do not use 5k results for the final reproducibility gate in Step 10.